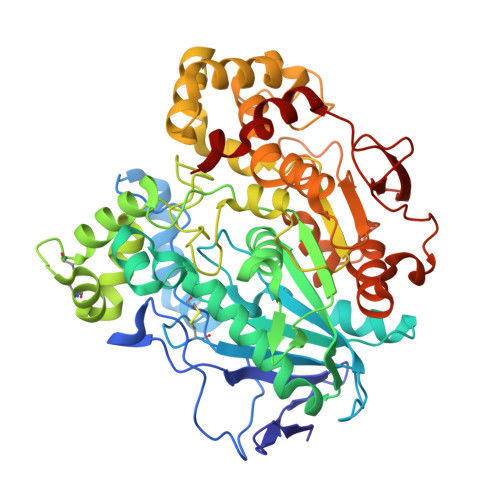
Three-dimensional structure of homodimeric cholesterol esterase-ligand complex at 1.4 A resolution.
Pletnev, V., Addlagatta, A., Wawrzak, Z., Duax, W.(2003) Acta Crystallogr D Biol Crystallogr 59: 50-56
- PubMed: 12499539
- DOI: https://doi.org/10.1107/s0907444902018851
- Primary Citation of Related Structures:
1LLF - PubMed Abstract:
The three-dimensional structure of a Candida cylindracea cholesterol esterase (ChE) homodimer (534 x 2 amino acids) in complex with a ligand of proposed formula C(23)H(48)O(2) has been determined at 1.4 A resolution in space group P1 using synchrotron low-temperature data. The structure refined to R = 0.136 and R(free) = 0.169 and has revealed new stereochemical details in addition to those detected for the apo- and holo-forms at 1.9 and 2.0 A resolution, respectively [Ghosh et al. (1995), Structure, 3, 279-288]. The cholesterol esterase structure is a dimer with four spatially separated interfacial contact areas and two symmetry-related pairs of openings to an internal intradimer cavity. Hydrophobic active-site gorges in each subunit face each other across a central interfacial cavity. The ChE subunits have carbohydrate chains attached to their Asn314 and Asn351 residues, with two ordered N-acetyl-D-glucosoamine moieties visible at each site. The side chains of 14 residues have two alternative conformations with occupancy values of 0.5 +/- 0.2. For each subunit the electron density in the enzyme active-site gorge is well modeled by a C(23)-chain fatty acid.
- Institute of Bioorganic Chemistry, Russian Academy of Science, Moscow, Russia.
Organizational Affiliation: