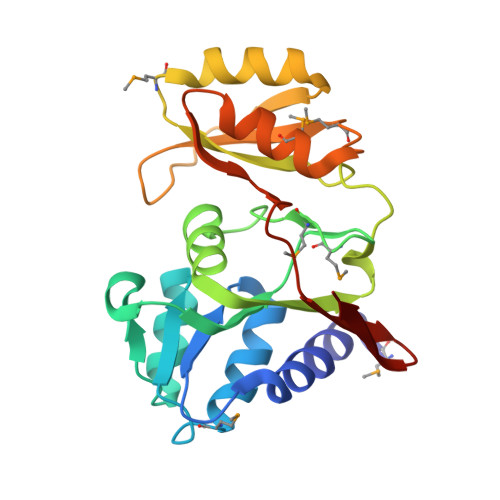
Crystal structure of D-ribose-5-phosphate isomerase (RpiA) from Escherichia coli
Rangarajan, E.S., Sivaraman, J., Matte, A., Cygler, M.(2002) Proteins 48: 737-740
- PubMed: 12211039 Search on PubMed
- DOI: https://doi.org/10.1002/prot.10203
- Primary Citation Related Structures:
1LKZ - Department of Biochemistry, McGill University, Montreal, Quebec, Canada.
Organizational Affiliation: