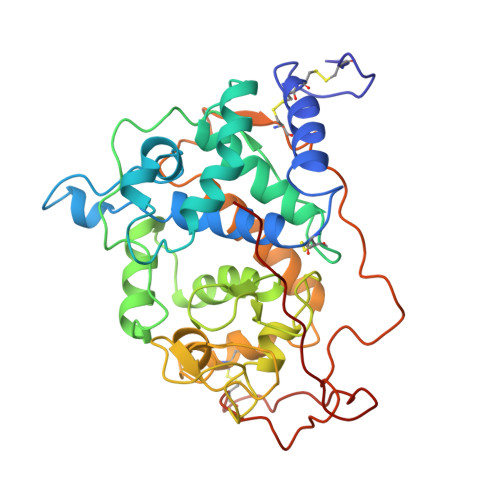
Crystallographic refinement of lignin peroxidase at 2 A.
Poulos, T.L., Edwards, S.L., Wariishi, H., Gold, M.H.(1993) J Biological Chem 268: 4429-4440
- PubMed: 8440725
- DOI: https://doi.org/10.2210/pdb1lga/pdb
- Primary Citation Related Structures:
1LGA - PubMed Abstract:
The crystal structure of the major lignin peroxidase isozyme from Phanerocheate chrysosporium has been refined to an R = 0.15 for data between 8 A and 2.03 A. The refined model consists of 2 lignin peroxidase molecules in the asymmetric unit, 2 calcium ions per monomer, 1 glucosamine per monomer N-linked to Asn-257, and 476 water molecules per asymmetric unit. The model exhibits excellent geometry with a root mean square deviation from ideality in bond distances and angles of 0.014 A and 2.9 degrees, respectively. Molecule 1 consists of all 343 residues, while molecule 2 consists of residues 1-341. The overall root mean square deviation in backbone atoms between the 2 molecules in the asymmetric unit is 0.36 A. The refinement at 2.0 A confirms our conclusions based on the partially refined 2.6-A structure (Edwards, S. L., Raag, R., Wariishi, H., Gold, M. H., and Poulos, T. L. (1993) Proc. Natl. Acad. Sci. U.S.A. 90, 750-754). The overall fold of lignin peroxidase closely resembles that of cytochrome c peroxidase. A superimposition of alpha-carbons gives a root mean square deviation of 2.65 A between the two peroxidases and 1.66 A for the helices. The active sites also are similar since both contain a proximal histidine heme ligand hydrogen-bonded to a buried aspartate residue and both contain histidine and arginine residues in the distal peroxide binding pocket. The most obvious difference in the active site is that whereas cytochrome c peroxidase has tryptophan residues located in the proximal and distal heme pockets, lignin peroxidase has phenylalanines. There are four other especially noteworthy differences in the two structures. First, although the heme in cytochrome c peroxidase is recessed about 10 A from the molecular surface, the heme pocket is open to solvent. The analogous opening in lignin peroxidase is smaller which can explain in part the differences in reactivity of the two hemes. This same opening may provide the site for binding small aromatic substrates. Second, lignin peroxidase has a carboxylate-carboxylate hydrogen bond important for heme binding that is not present in cytochrome c peroxidase. Third, lignin peroxidase contains 2 structural calcium ions while cytochrome c peroxidase contains no calcium. The calciums in lignin peroxidase coordinate to residues near the C-terminal ends of the distal and proximal helices and hence are probably important for maintaining the integrity of the active site. Fourth, the extra 49 residues in lignin peroxidase not present in cytochrome c peroxidase constitutes the C-terminal end of the molecule with the C terminus situated at the "front" end of the molecule between the two heme propionates.
- Department of Molecular Biology, University of California, Irvine 92717.
Organizational Affiliation: