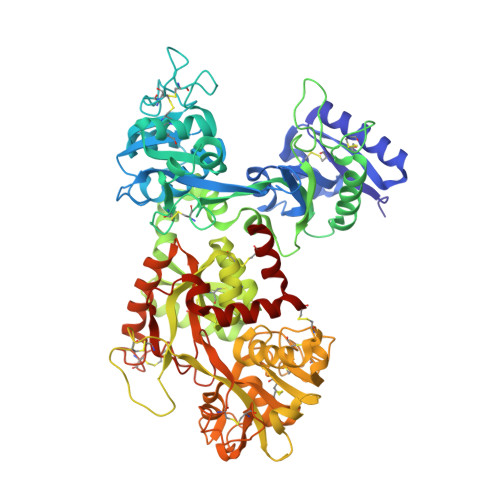
Molecular replacement solution of the structure of apolactoferrin, a protein displaying large-scale conformational change.
Norris, G.E., Anderson, B.F., Baker, E.N.(1991) Acta Crystallogr B 47: 998-1004
- PubMed: 1772635
- DOI: https://doi.org/10.1107/s0108768191008418
- Primary Citation of Related Structures:
1LFH - PubMed Abstract:
The crystal structure of an orthorhombic form of human apolactoferrin (ApoLf) has been determined from 2.8 A diffractometer data by molecular replacement methods. A variety of search models derived from the diferric lactoferrin structure (Fe2Lf) were used to obtain a consistent solution to the rotation function. An R-factor search gave the correct translational solution and the model was refined by rigid-body least-squares refinement (program CORELS). Only three of the four domains were located correctly by this procedure, however; the fourth was finally placed correctly by rotating it manually onto three strands of electron density which were recognized as part of its central beta-sheet. The final model, refined by restrained least-squares methods to an R factor of 0.214 for data in the resolution range 10.0 to 2.8 A, shows a large domain movement in the N-terminal half of the molecule (a 54 degree rotation of domain N2) and smaller domain movements elsewhere, when compared with Fe2Lf. A feature of the crystal structure is that although the ApoLf and Fe2Lf unit cells appear very similar, their crystal packing and molecular structures are quite different.
- Department of Chemistry and Biochemistry, Massey University, Palmerston North, New Zealand.
Organizational Affiliation: