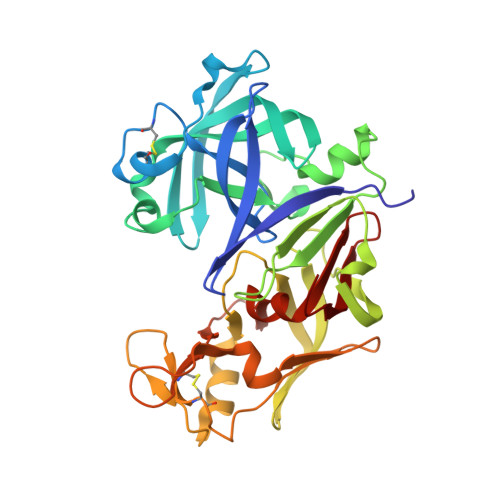
Structures of Ser205 mutant plasmepsin II from Plasmodium falciparum at 1.8 A in complex with the inhibitors rs367 and rs370.
Asojo, O.A., Afonina, E., Gulnik, S.V., Yu, B., Erickson, J.W., Randad, R., Medjahed, D., Silva, A.M.(2002) Acta Crystallogr D Biol Crystallogr 58: 2001-2008
- PubMed: 12454457 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444902014695
- Primary Citation Related Structures:
1LEE, 1LF2 - PubMed Abstract:
Plasmepsin II is one of the four catalytically active plasmepsins found in the food vacuole of Plasmodium falciparum. These enzymes initiate hemoglobin degradation by cleavage at the alpha-chain between Phe33 and Leu34. The crystal structures of Ser205 mutant plasmepsin II from P. falciparum in complex with two inhibitors have been refined at a resolution of 1.8 A in the space group I222 and to R factors of 19.9 and 19.5%. Each crystal contains one monomer in the asymmetric unit. Both inhibitors have a Phe-Leu core and incorporate tetrahedral transition-state mimetic hydroxypropylamine. The inhibitor rs367 possesses a 2,6-dimethylphenyloxyacetyl group at the P2 position and 3-aminobenzamide at the P2' position, while rs370 has the same P2 group but 4-aminobenzamide in the P2' position. These complexes reveal key conserved hydrogen bonds between the inhibitor and the binding-cavity residues, notably with the flap residues Val78 and Ser79, the catalytic dyad Asp34 and Asp214 and the residues Ser218 and Gly36 that are in proximity to the catalytic dyad. The structures also show unexpected conformational variability of the binding cavity of plasmepsin II and may reflect the mode of binding of the hemoglobin alpha-chain for cleavage.
- Structure Biochemistry Program, National Cancer Institute/SAIC, Frederick, MD 21702, USA. asojo@ncifcrf.gov
Organizational Affiliation: