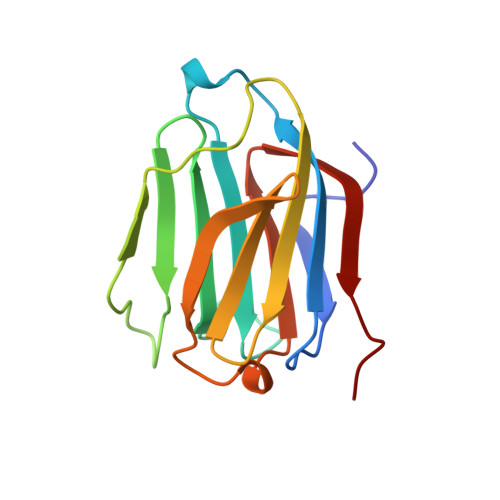
Crystal structure of human Charcot-Leyden crystal protein, an eosinophil lysophospholipase, identifies it as a new member of the carbohydrate-binding family of galectins.
Leonidas, D.D., Elbert, B.L., Zhou, Z., Leffler, H., Ackerman, S.J., Acharya, K.R.(1995) Structure 3: 1379-1393
- PubMed: 8747464 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00275-1
- Primary Citation Related Structures:
1LCL - PubMed Abstract:
The Charcot-Leyden crystal (CLC) protein is a major autocrystallizing constituent of human eosinophils and basophils, comprising approximately 10% of the total cellular protein in these granulocytes. Identification of the distinctive hexagonal bipyramidal crystals of CLC protein in body fluids and secretions has long been considered a hallmark of eosinophil-associated allergic inflammation. Although CLC protein possesses lysophospholipase activity, its role(s) in eosinophil or basophil function or associated inflammatory responses has remained speculative. The crystal structure of the CLC protein has been determined at 1.8 A resolution using X-ray crystallography. The overall structural fold of CLC protein is highly similar to that of galectins -1 and -2, members of an animal lectin family formerly classified as S-type or S-Lac (soluble lactose-binding) lectins. This is the first structure of an eosinophil protein to be determined and the highest resolution structure so far determined for any member of the galectin family. The CLC protein structure possesses a carbohydrate-recognition domain comprising most, but not all, of the carbohydrate-binding residues that are conserved among the galectins. The protein exhibits specific (albeit weak) carbohydrate-binding activity for simple saccharides including N-acetyl-D-glucosamine and lactose. Despite CLC protein having no significant sequence or structural similarities to other lysophospholipase catalytic triad has also been identified within the CLC structure, making it a unique dual-function polypeptide. These structural findings suggest a potential intracellular and/or extracellular role(s) for the galectin-associated activities of CLC protein in eosinophil and basophil function in allergic diseases and inflammation.
- School of Biology and Biochemistry, University of Bath, Claverton Down, UK.
Organizational Affiliation: