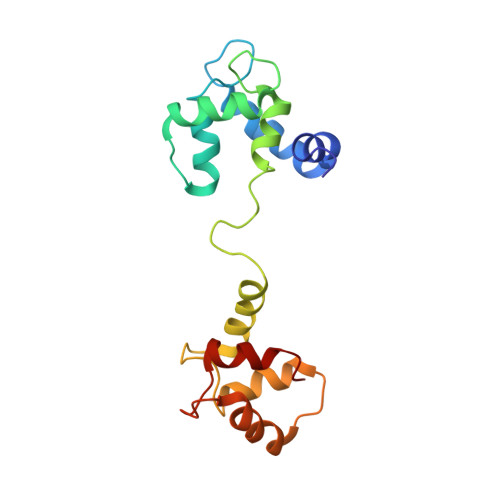
Solution Structure of Calcium-Saturated Cardiac Troponin C bound to cardiac Troponin I.
Dvoretsky, A., Abusamhadneh, E.M., Howarth, J.W., Rosevear, P.R.(2002) J Biological Chem 277: 38565-38570
- PubMed: 12147696
- DOI: https://doi.org/10.1074/jbc.M205306200
- Primary Citation Related Structures:
1LA0 - PubMed Abstract:
Cardiac troponin C (TnC) is composed of two globular domains connected by a flexible linker. In solution, linker flexibility results in an ill defined orientation of the two globular domains relative to one another. We have previously shown a decrease in linker flexibility in response to cardiac troponin I (cTnI) binding. To investigate the relative orientation of calcium-saturated TnC domains when bound to cTnI, (1)H-(15)N residual dipolar couplings were measured in two different alignment media. Similarity in alignment tensor orientation for the two TnC domains supports restriction of domain motion in the presence of cTnI. The relative spatial orientation of TnC domains bound to TnI was calculated from measured residual dipolar couplings and long-range distance restraints utilizing a rigid body molecular dynamics protocol. The relative domain orientation is such that hydrophobic pockets face each other, forming a latch to constrain separate helical segments of TnI. We have utilized this structure to successfully explain the observed functional consequences of linker region deletion mutants. Together, these studies suggest that, although linker plasticity is important, the ability of TnC to function in muscle contraction can be correlated with a preferred domain orientation and interdomain distance.
- Department of Molecular Genetics, Biochemistry, and Microbiology, University of Cincinnati, College of Medicine, Cincinnati, Ohio 45267, USA.
Organizational Affiliation: