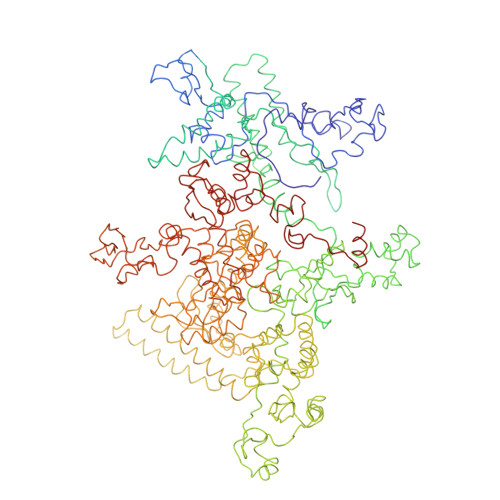
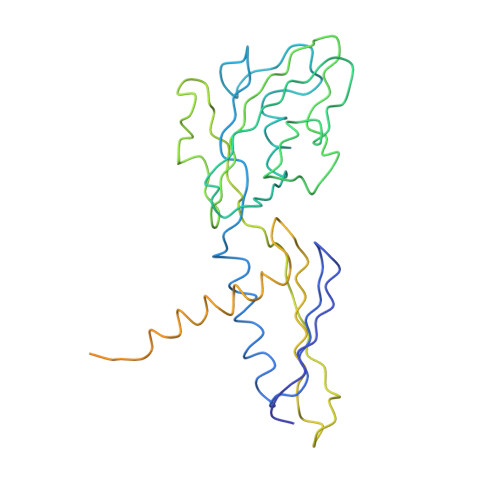
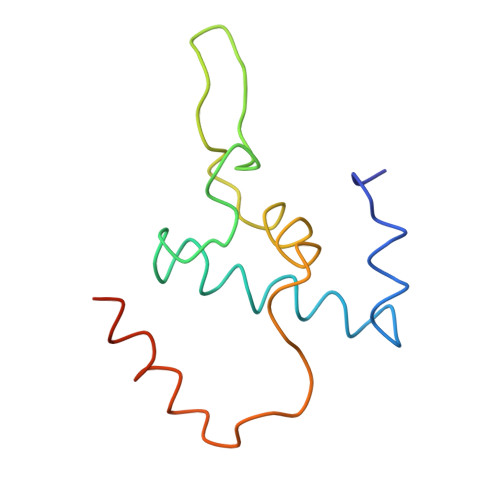
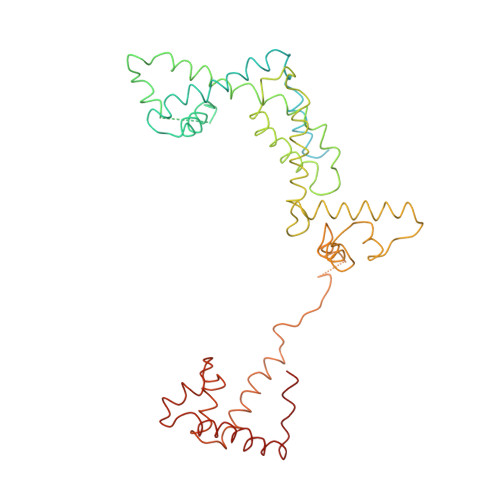
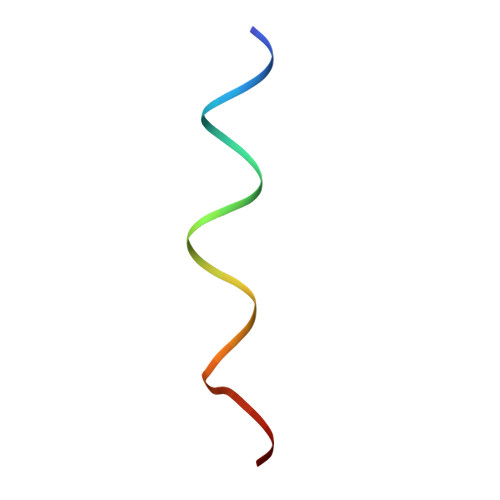Structural basis of transcription initiation: an RNA polymerase holoenzyme-DNA complex.
Murakami, K.S., Masuda, S., Campbell, E.A., Muzzin, O., Darst, S.A.(2002) Science 296: 1285-1290
- PubMed: 12016307
- DOI: https://doi.org/10.1126/science.1069595
- Primary Citation Related Structures:
1L9Z - PubMed Abstract:
The crystal structure of Thermus aquaticus RNA polymerase holoenzyme (alpha2betabeta'omegasigmaA) complexed with a fork-junction promoter DNA fragment has been determined by fitting high-resolution x-ray structures of individual components into a 6.5-angstrom resolution map. The DNA lies across one face of the holoenzyme, completely outside the RNA polymerase active site channel. All sequence-specific contacts with core promoter elements are mediated by the sigma subunit. A universally conserved tryptophan is ideally positioned to stack on the exposed face of the base pair at the upstream edge of the transcription bubble. Universally conserved basic residues of the sigma subunit provide critical contacts with the DNA phosphate backbone and play a role in directing the melted DNA template strand into the RNA polymerase active site. The structure explains how holoenzyme recognizes promoters containing variably spaced -10 and -35 elements and provides the basis for models of the closed and open promoter complexes.
- The Rockefeller University, 1230 York Avenue, New York, NY 10021, USA.
Organizational Affiliation: