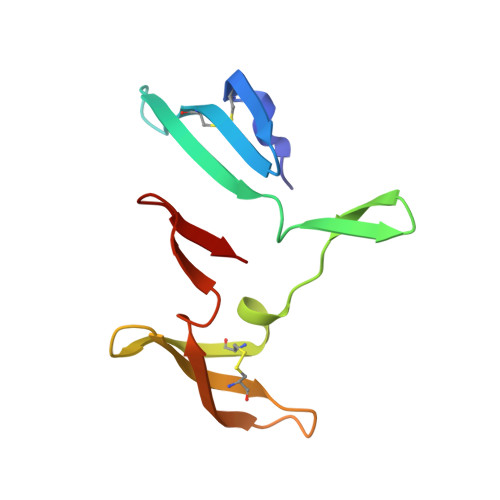
The domain-swapped dimer of cyanovirin-N is in a metastable folded state: reconciliation of X-ray and NMR structures.
Barrientos, L.G., Louis, J.M., Botos, I., Mori, T., Han, Z., O'Keefe, B.R., Boyd, M.R., Wlodawer, A., Gronenborn, A.M.(2002) Structure 10: 673-686
- PubMed: 12015150 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00758-x
- Primary Citation Related Structures:
1L5B, 1L5E - PubMed Abstract:
The structure of the potent HIV-inactivating protein cyanovirin-N was previously found by NMR to be a monomer in solution and a domain-swapped dimer by X-ray crystallography. Here we demonstrate that, in solution, CV-N can exist both in monomeric and in domain-swapped dimeric form. The dimer is a metastable, kinetically trapped structure at neutral pH and room temperature. Based on orientational NMR constraints, we show that the domain-swapped solution dimer is similar to structures in two different crystal forms, exhibiting solely a small reorientation around the hinge region. Mutation of the single proline residue in the hinge to glycine significantly stabilizes the protein in both its monomeric and dimeric forms. By contrast, mutation of the neighboring serine to proline results in an exclusively dimeric protein, caused by a drastic destabilization of the monomer.
- Laboratory of Chemical Physics, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, MD 20892, USA.
Organizational Affiliation: