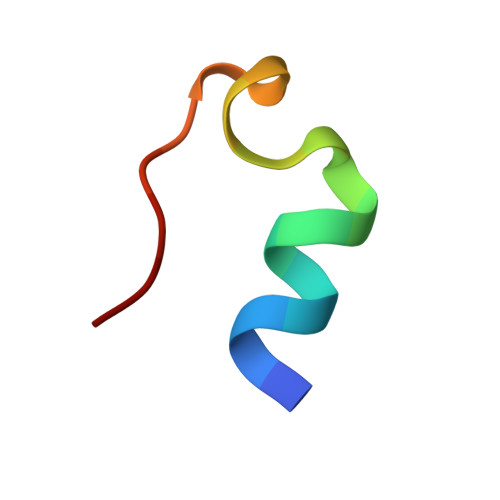
Designing a 20-residue protein.
Neidigh, J.W., Fesinmeyer, R.M., Andersen, N.H.(2002) Nat Struct Biol 9: 425-430
- PubMed: 11979279 Search on PubMed
- DOI: https://doi.org/10.1038/nsb798
- Primary Citation Related Structures:
1L2Y - PubMed Abstract:
Truncation and mutation of a poorly folded 39-residue peptide has produced 20-residue constructs that are >95% folded in water at physiological pH. These constructs optimize a novel fold, designated as the 'Trp-cage' motif, and are significantly more stable than any other miniprotein reported to date. Folding is cooperative and hydrophobically driven by the encapsulation of a Trp side chain in a sheath of Pro rings. As the smallest protein-like construct, Trp-cage miniproteins should provide a testing ground for both experimental studies and computational simulations of protein folding and unfolding pathways. Pro Trp interactions may be a particularly effective strategy for the a priori design of self-folding peptides.
- Department of Chemistry, University of Washington, Seattle, Washington 98195, USA.
Organizational Affiliation: