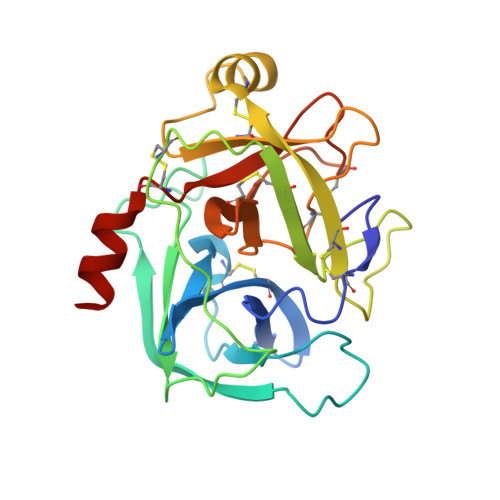
Crystal structure and biochemical characterization of human kallikrein 6 reveals that a trypsin-like kallikrein is expressed in the central nervous system.
Bernett, M.J., Blaber, S.I., Scarisbrick, I.A., Dhanarajan, P., Thompson, S.M., Blaber, M.(2002) J Biological Chem 277: 24562-24570
- PubMed: 11983703 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M202392200
- Primary Citation Related Structures:
1L2E - PubMed Abstract:
The human kallikreins are a large multigene family of closely related serine-type proteases. In this regard, they are similar to the multigene kallikrein families characterized in mice and rats. There is a much more extensive body of knowledge regarding the function of mouse and rat kallikreins in comparison with the human kallikreins. Human kallikrein 6 has been proposed as the homologue to rat myelencephalon-specific protease, an arginine-specific degradative-type protease abundantly expressed in the central nervous system and implicated in demyelinating disease. We present the x-ray crystal structure of mature, active recombinant human kallikrein 6 at 1.75-A resolution. This high resolution model provides the first three-dimensional view of one of the human kallikreins and one of only a few structures of serine proteases predominantly expressed in the central nervous system. Enzymatic data are presented that support the identification of human kallikrein 6 as the functional homologue of rat myelencephalon-specific protease and are corroborated by a molecular phylogenetic analysis. Furthermore, the x-ray data provide support for the characterization of human kallikrein 6 as a degradative protease with structural features more similar to trypsin than the regulatory kallikreins.
- Institute of Molecular Biophysics, Department of Chemistry and Biochemistry, Florida State University, Tallahassee, Florida 32306-4380, USA.
Organizational Affiliation: