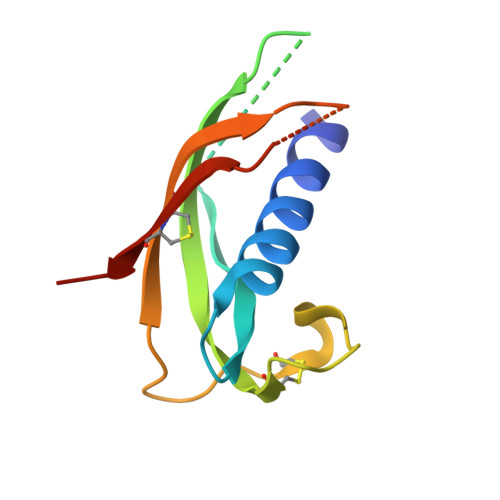
Structure of the cathelicidin motif of protegrin-3 precursor: structural insights into the activation mechanism of an antimicrobial protein.
Sanchez, J.F., Hoh, F., Strub, M.P., Aumelas, A., Dumas, C.(2002) Structure 10: 1363-1370
- PubMed: 12377122
- DOI: https://doi.org/10.1016/s0969-2126(02)00859-6
- Primary Citation Related Structures:
1KWI, 1LXE - PubMed Abstract:
Cathelicidins are a family of antimicrobial proteins isolated from leucocytes and epithelia cells that contribute to the innate host defense mechanisms in mammalians. Located in the C-terminal part of the holoprotein, the cathelicidin-derived antimicrobial peptide is liberated by a specific protease cleavage. Here, we report the X-ray structure of the cathelicidin motif of protegrin-3 solved by MAD phasing using the selenocysteine-labeled protein. Its overall structure represents a fold homologous to the cystatin family and adopts two native states, a monomer, and a domain-swapped dimer. This crystal structure is the first example of a structural characterization of the highly conserved cathelicidin motif and thus provides insights into the possible mechanism of activation of the antimicrobial protegrin peptide.
- Centre de Biochimie Structurale, UMR CNRS 5048, UMR 554 INSERM, Université Montpellier I, 15 Avenue Charles Flahault, 34060 Montpellier Cedex, France.
Organizational Affiliation: