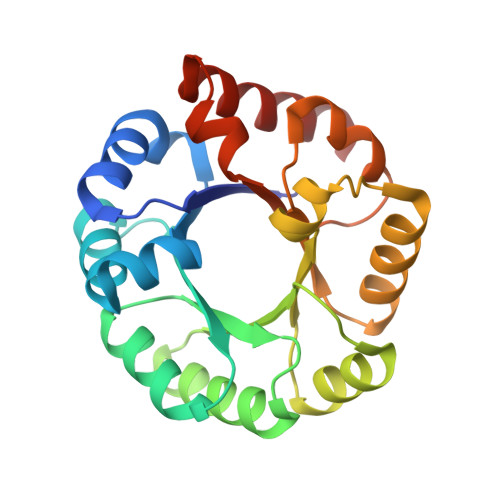
Homologous (beta/alpha)8-barrel enzymes that catalyze unrelated reactions: orotidine 5'-monophosphate decarboxylase and 3-keto-L-gulonate 6-phosphate decarboxylase.
Wise, E., Yew, W.S., Babbitt, P.C., Gerlt, J.A., Rayment, I.(2002) Biochemistry 41: 3861-3869
- PubMed: 11900527
- DOI: https://doi.org/10.1021/bi012174e
- Primary Citation Related Structures:
1KV8, 1KW1 - PubMed Abstract:
The 3-keto-L-gulonate 6-phosphate decarboxylase (KGPDC) encoded by the ulaD gene in the Escherichia coli genome [Yew, W. S., and Gerlt, J. A. (2002) J. Bacteriol. 184, 302-306] and orotidine 5'-monophosphate decarboxylase (OMPDC) are homologous (derived from a common ancestor) but catalyze different reactions. The metal-independent decarboxylation reaction catalyzed by OMPDC avoids the formation of a vinyl anion intermediate; the Mg2+-dependent decarboxylation reaction catalyzed by KGPDC involves the formation of an enediolate anion intermediate. Based on the available structures of OMPDC, a sequence alignment allows the predictions that (1) KGPDC is a dimer of (beta/alpha)8-barrels, with the active sites located at the dimer interface; (2) KGPDC and OMPDC share an aspartate residue at the end of the first beta-strand and an Asp-x-Lys-x-x-Asp motif at the end of the third beta-strand with OMPDC; but (3) KGPDC has a Glu instead of a Lys at the end of the second beta-strand. The structure of KGPDC has been determined in the presence of Mg2+ and the substrate analogue L-gulonate 6-phosphate and confirms these predictions. The carboxylate functional groups at the ends of the first, second, and third beta-strands in KGPDC are ligands of the Mg2+; in OMPDC, the homologues of these residues participate in a hydrogen-bonded network that facilitates the decarboxylation reaction. The 3-OH group of the substrate analogue is coordinated to the Mg2+, supporting the hypothesis that the mechanism of the decarboxylation catalyzed by KGPDC involves stabilization of an enediolate anion intermediate. These structural studies establish the existence of the OMPDC "suprafamily," in which members catalyze reactions that occur in different metabolic pathways and share no mechanistic relationship. The existence of this suprafamily demonstrates that divergent evolution can be opportunistic, conscripting active site features of a progenitor to catalyze unrelated functions. Accordingly, sequence or structure homology alone cannot be used to infer the functions of new proteins discovered in genome projects.
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53706, USA.
Organizational Affiliation: