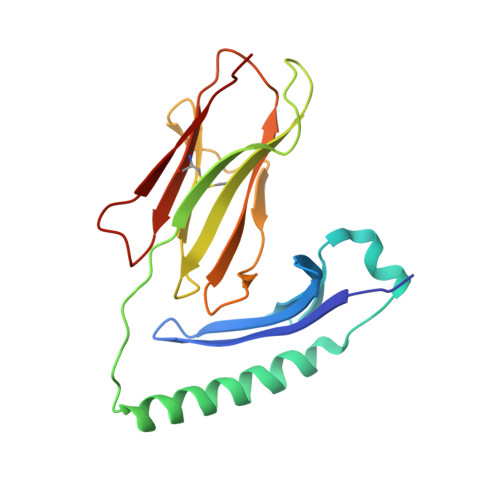
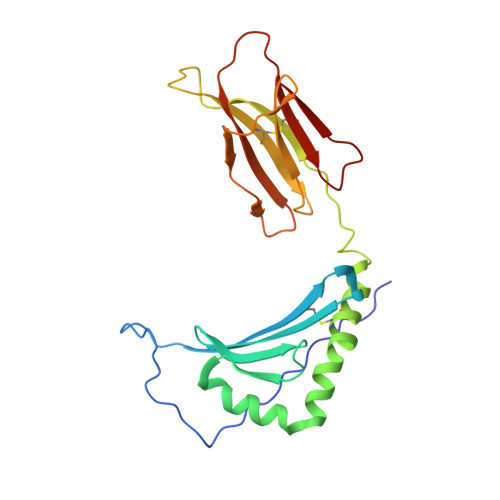
Structural basis of cytochrome c presentation by IE(k).
Fremont, D.H., Dai, S., Chiang, H., Crawford, F., Marrack, P., Kappler, J.(2002) J Exp Medicine 195: 1043-1052
- PubMed: 11956295 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1084/jem.20011971
- Primary Citation Related Structures:
1KT2, 1KTD - PubMed Abstract:
The COOH-terminal peptides of pigeon and moth cytochrome c, bound to mouse IE(k), are two of the most thoroughly studied T cell antigens. We have solved the crystal structures of the moth peptide and a weak agonist-antagonist variant of the pigeon peptide bound to IE(k). The moth peptide and all other peptides whose structures have been solved bound to IE(k), have a lysine filling the p9 pocket of IE(k). However, the pigeon peptide has an alanine at p9 shifting the lysine to p10. Rather than kinking to place the lysine in the anchor pocket, the pigeon peptide takes the extended course through the binding groove, which is characteristic of all other peptides bound to major histocompatibility complex (MHC) class II. Thus, unlike MHC class I, in which peptides often kink to place optimally anchoring side chains, MHC class II imposes an extended peptide conformation even at the cost of a highly conserved anchor residue. The substitution of Ser for Thr at p8 in the variant pigeon peptide induces no detectable surface change other than the loss of the side chain methyl group, despite the dramatic change in recognition by T cells. Finally, these structures can be used to interpret the many published mutational studies of these ligands and the T cell receptors that recognize them.
- Department of Pathology and Immunology, Washington University School of Medicine, St Louis, MO 63110, USA.
Organizational Affiliation: