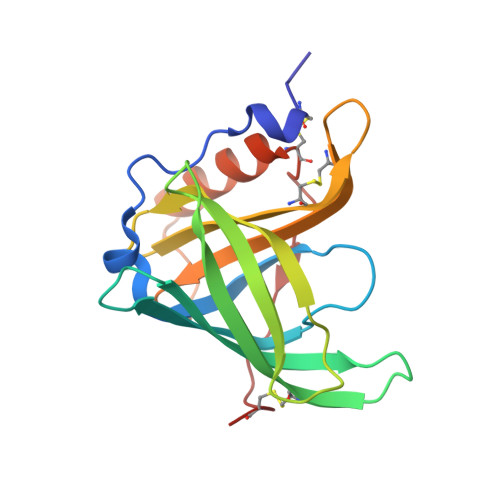
High-resolution Structures of Retinol-binding Protein in Complex with Retinol: pH-induced Protein Structural Changes in the Crystal State
Calderone, V., Berni, R., Zanotti, G.(2003) J Mol Biology 329: 841-850
- PubMed: 12787682 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00468-6
- Primary Citation Related Structures:
1KT3, 1KT4, 1KT5, 1KT6, 1KT7 - PubMed Abstract:
The targeted delivery of non-polar ligands by binding proteins to membranes or membrane receptors involves the release of these ligands on or near the plasma membrane of target cells. Because these hydrophobic ligands are often bound inside a deep cavity of binding proteins, as shown previously for plasma retinol-binding protein (RBP), their release from these proteins might require the destabilization of the protein structure by partially denaturing conditions, such as those possibly present near plasma membranes. RBP is a plasma transport protein which delivers specifically retinol from its store sites to target cells. Here, we report the high-resolution (1.1-1.4A) crystal structures of bovine holo-RBP at five different pH values, ranging from 9 to 2. While unraveling details of the native protein structure and of the interactions with retinol at nearly atomic resolution at neutral pH, this study provides evidence for definite pH-induced modifications of several structural features of RBP. The structure most representative of the changes that holo-RBP undergoes at different pH values is that of its flexible state at pH 2. At this pH, most significant are the alteration of the arrangement of salt bridges and of the network of water molecules/H-bonds that participates in the retinol-RBP interaction, an appreciable increase of the volume of the beta-barrel cavity, a considerably higher degree of mobility of the RBP-bound ligand and of several protein regions and the disorder of a large number of solvent molecules that are ordered at neutral pH. These changes are likely to be accompanied by a modification of the pattern of charge distribution on the protein surface. All these changes, which reveal a substantially lowered conformational stability of RBP, presumably occur at the initial stages of the acidic denaturation of RBP and are possibly associated with a facilitated release of the retinol molecule from its carrier protein.
- Department of Organic Chemistry, University of Padova, and Biopolymer Research Center, Italian National Research Council, Via Marzolo 1, 35131, Padova, Italy.
Organizational Affiliation: