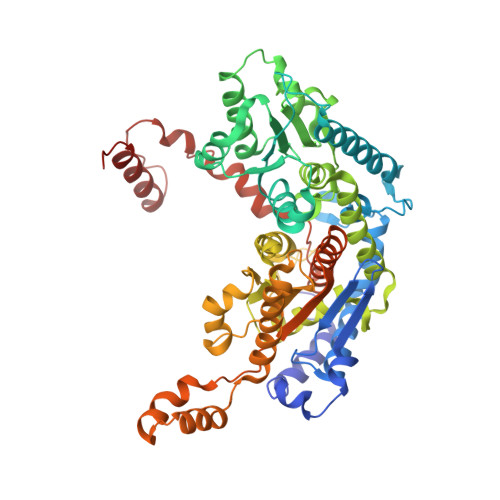
The crystal structure of rabbit phosphoglucose isomerase complexed with 5-phospho-D-arabinonohydroxamic acid.
Arsenieva, D., Hardre, R., Salmon, L., Jeffery, C.J.(2002) Proc Natl Acad Sci U S A 99: 5872-5877
- PubMed: 11983887
- DOI: https://doi.org/10.1073/pnas.052131799
- Primary Citation of Related Structures:
1KOJ - PubMed Abstract:
Phosphoglucose isomerase (EC ) catalyzes the second step in glycolysis, the reversible isomerization of D-glucose 6-phosphate to D-fructose 6-phosphate. The reaction mechanism involves acid-base catalysis with proton transfer and proceeds through a cis-enediol(ate) intermediate. 5-Phospho-D-arabinonohydroxamic acid (5PAH) is a synthetic small molecule that resembles the reaction intermediate, differing only in that it has a nitrogen atom in place of C1. Hence, 5PAH is the best inhibitor of the isomerization reaction reported to date with a K(i) of 2 x 10(-7) M. Here we report the crystal structure of rabbit phosphoglucose isomerase complexed with 5PAH at 1.9 A resolution. The interaction of 5PAH with amino acid residues in the enzyme active site supports a model of the catalytic mechanism in which Glu-357 transfers a proton between C1 and C2 and Arg-272 helps stabilize the intermediate. It also suggests a mechanism for proton transfer between O1 and O2.
- Laboratory for Molecular Biology, MC567, Department of Biological Sciences, University of Illinois, Chicago, IL 60607, USA.
Organizational Affiliation: