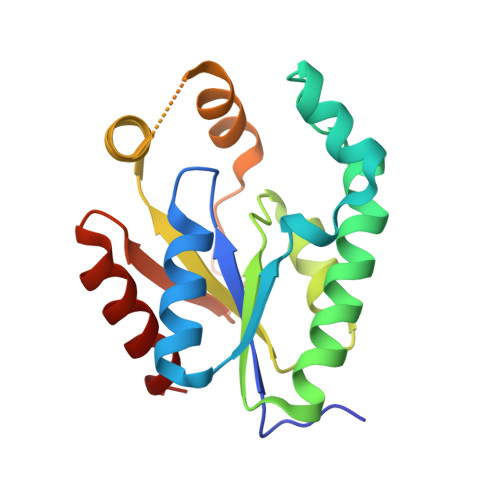
Conformational changes during the catalytic cycle of gluconate kinase as revealed by X-ray crystallography.
Kraft, L., Sprenger, G.A., Lindqvist, Y.(2002) J Mol Biology 318: 1057-1069
- PubMed: 12054802 Search on PubMed
- DOI: https://doi.org/10.1016/S0022-2836(02)00215-2
- Primary Citation Related Structures:
1KNQ, 1KO1, 1KO4, 1KO5, 1KO8, 1KOF - PubMed Abstract:
The crystal structure of gluconate kinase from Escherichia coli has been determined to 2.0 A resolution by X-ray crystallography. The three-dimensional structure was solved by multi-wavelength anomalous dispersion, using a crystal of selenomethionine-substituted enzyme. Gluconate kinase is an alpha/beta structure consisting of a twisted parallel beta-sheet surrounded by alpha-helices with overall topology similar to nucleoside monophosphate (NMP) kinases, such as adenylate kinase. In order to identify residues involved in substrate binding and catalysis, structures of binary complexes with ATP, the ATP analogue adenosine 5'-(beta,gamma-methylene) triphosphate and the product, gluconate-6-phosphate have been determined. Significant conformational changes are induced upon binding of ATP to the enzyme. The largest changes involve a hinge-bending motion of the NMP(bind) part and a motion of the LID with adjacent helices, which opens the cavity to the second substrate, gluconate. Opening of the active site cleft upon ATP binding is the opposite of what has been observed in the NMP kinase family so far, which usually close their active site to prevent fortuitous hydrolysis of ATP. The conformational change positions the side-chain of Arg120 to stack with the purine ring of ATP and the side-chain of Arg124 is shifted to interact with the alpha-phosphate in ATP, at the same time protecting ATP from solvent water. The beta and gamma-phosphate groups of ATP bind in the predicted P-loop. A conserved lysine side-chain interacts with the gamma-phosphate group, and might promote phosphoryl transfer. Gluconate-6-phosphate binds with its phosphate group in a similar position as the gamma-phosphate of ATP, consistent with inline phosphoryl transfer. The gluconate binding-pocket in GntK is located in a different position than the nucleoside binding-site usually found in NMP kinases.
- Molecular Structural Biology, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, SE-17177 Stockholm, Sweden.
Organizational Affiliation: