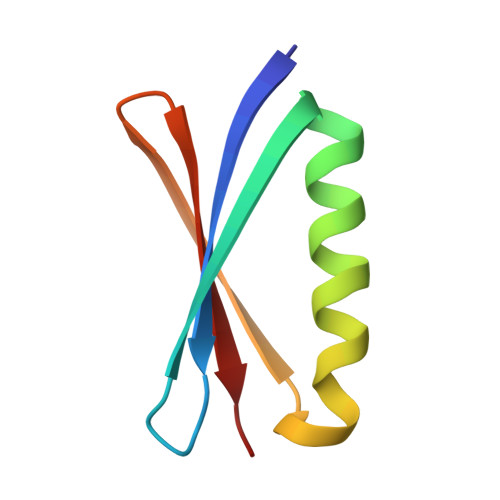
Accurate computer-based design of a new backbone conformation in the second turn of protein L.
Kuhlman, B., O'Neill, J.W., Kim, D.E., Zhang, K.Y., Baker, D.(2002) J Mol Biology 315: 471-477
- PubMed: 11786026 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.5229
- Primary Citation Related Structures:
1KH0 - PubMed Abstract:
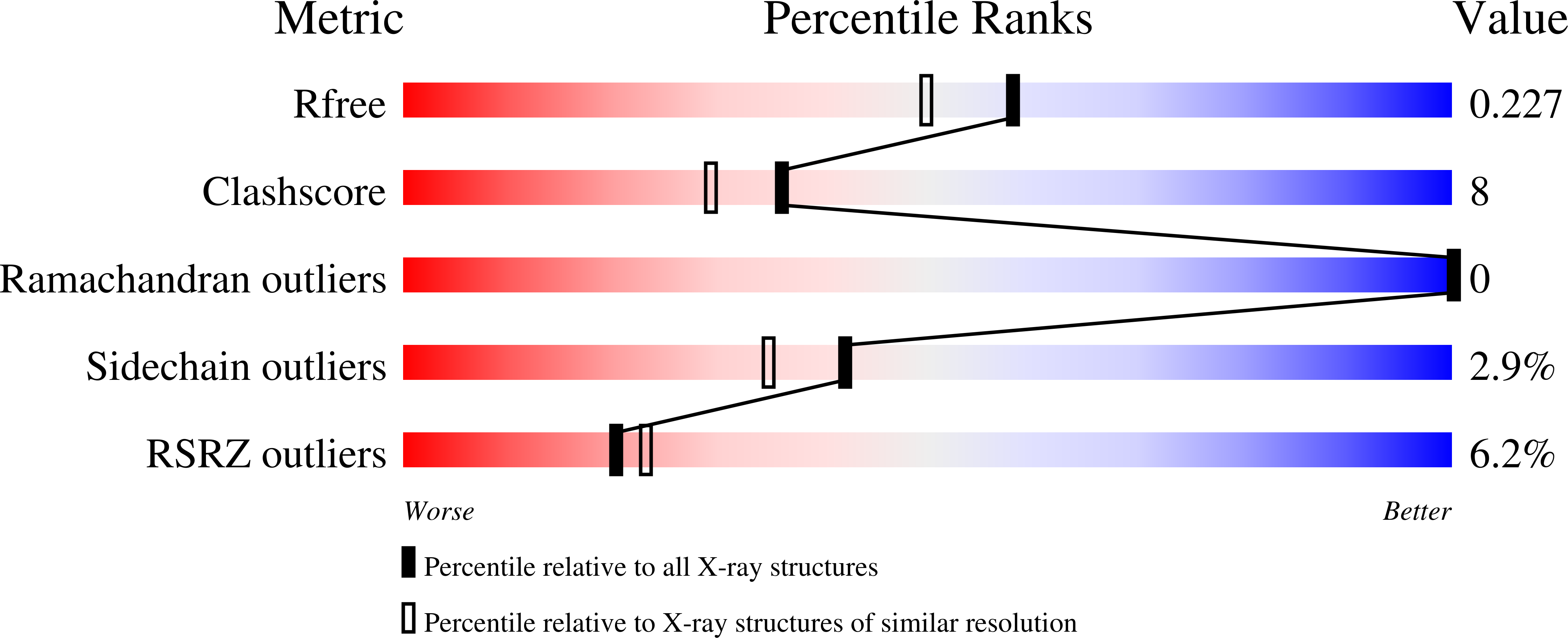
The rational design of loops and turns is a key step towards creating proteins with new functions. We used a computational design procedure to create new backbone conformations in the second turn of protein L. The Protein Data Bank was searched for alternative turn conformations, and sequences optimal for these turns in the context of protein L were identified using a Monte Carlo search procedure and an energy function that favors close packing. Two variants containing 12 and 14 mutations were found to be as stable as wild-type protein L. The crystal structure of one of the variants has been solved at a resolution of 1.9 A, and the backbone conformation in the second turn is remarkably close to that of the in silico model (1.1 A RMSD) while it differs significantly from that of wild-type protein L (the turn residues are displaced by an average of 7.2 A). The folding rates of the redesigned proteins are greater than that of the wild-type protein and in contrast to wild-type protein L the second beta-turn appears to be formed at the rate limiting step in folding.
- Department of Biochemistry, University of Washington, Seattle, WA 98195, USA.
Organizational Affiliation: