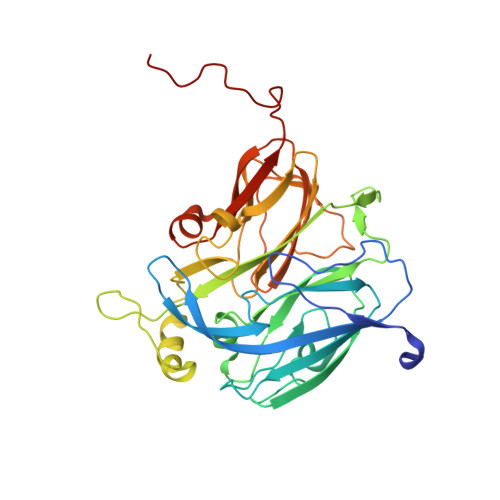
Crystal structure of a NO-forming nitrite reductase mutant: an analog of a transition state in enzymatic reaction
Liu, S.Q., Chang, T., Liu, M.Y., LeGall, J., Chang, W.C., Zhang, J.P., Liang, D.C., Chang, W.R.(2003) Biochem Biophys Res Commun 302: 568-574
- PubMed: 12615072 Search on PubMed
- DOI: https://doi.org/10.1016/s0006-291x(03)00166-9
- Primary Citation Related Structures:
1KCB - PubMed Abstract:
I257E was obtained by site directed mutagenesis of nitrite reductase from Achromobacter cycloclastes. The mutant has no enzyme activity. Its crystal structure determined at 1.65A resolution shows that the side-chain carboxyl group of the mutated residue, Glu257, coordinates with the type 2 copper in the mutant and blocks the contact between the type 2 copper and its solvent channel, indicating that the accessibility of the type 2 copper is essential for maintaining the activity of nitrite reductase. The carboxylate is an analog of the substrate, nitrite, but the distances between the type 2 copper and the two oxygen atoms of the side-chain carboxyl group are reversed in comparison to the binding of nitrite to the native enzyme. In the mutant, both the type 2 copper and the N epsilon atom on the imidazole ring of its coordinated residue His135 move in the substrate binding direction relative to the native enzyme. In addition, an EPR study showed that the type 2 copper in the mutant is in a reduced state. We propose that mutant I257E is in a state corresponding to a transition state in the enzymatic reaction.
- National laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Science, Beijing 100101, China.
Organizational Affiliation: