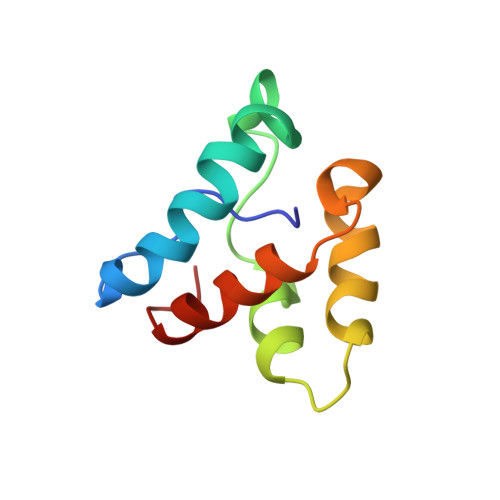
Solution NMR Structure of the Myosin Phosphatase Inhibitor Protein CPI-17 Shows Phosphorylation-induced Conformational Changes Responsible for Activation
Ohki, S., Eto, M., Kariya, E., Hayano, T., Hayashi, Y., Yazawa, M., Brautigan, D., Kainosho, M.(2001) J Mol Biology 314: 839-849
- PubMed: 11734001
- DOI: https://doi.org/10.1006/jmbi.2001.5200
- Primary Citation of Related Structures:
1K5O - PubMed Abstract:
Contractility of vascular smooth muscle depends on phosphorylation of myosin light chains, and is modulated by hormonal control of myosin phosphatase activity. Signaling pathways activate kinases such as PKC or Rho-dependent kinases that phosphorylate the myosin phosphatase inhibitor protein called CPI-17. Phosphorylation of CPI-17 at Thr38 enhances its inhibitory potency 1000-fold, creating a molecular on/off switch for regulating contraction. We report the solution NMR structure of the CPI-17 inhibitory domain (residues 35-120), which retains the signature biological properties of the full-length protein. The final ensemble of 20 sets of NMR coordinates overlaid onto their mean structure with r.m.s.d. values of 0.84(+/-0.22) A for the backbone atoms. The protein forms a novel four-helix, V-shaped bundle comprised of a central anti-parallel helix pair (B/C helices) flanked by two large spiral loops formed by the N and C termini that are held together by another anti-parallel helix pair (A/D helices) stabilized by intercalated aromatic and aliphatic side-chains. Chemical shift perturbations indicated that phosphorylation of Thr38 induces a conformational change involving displacement of helix A, without significant movement of the other three helices. This conformational change seems to flex one arm of the molecule, thereby exposing new surfaces of the helix A and the nearby phosphorylation loop to form specific interactions with the catalytic site of the phosphatase. This phosphorylation-dependent conformational change offers new structural insights toward understanding the specificity of CPI-17 for myosin phosphatase and its function as a molecular switch.
- JST and Graduate School of Science, Tokyo Metropolitan University, 1-1 Minami-ohsawa, Tokyo, Hachioji, 192-0397, Japan.
Organizational Affiliation: