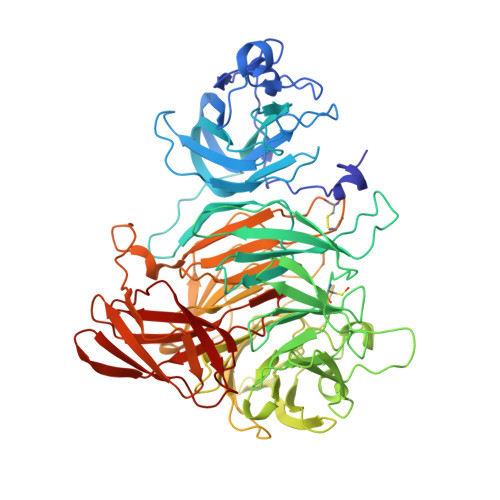
Crystal structure of the precursor of galactose oxidase: an unusual self-processing enzyme.
Firbank, S.J., Rogers, M.S., Wilmot, C.M., Dooley, D.M., Halcrow, M.A., Knowles, P.F., McPherson, M.J., Phillips, S.E.(2001) Proc Natl Acad Sci U S A 98: 12932-12937
- PubMed: 11698678 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.231463798
- Primary Citation Related Structures:
1K3I - PubMed Abstract:
Galactose oxidase (EC ) is a monomeric enzyme that contains a single copper ion and catalyses the stereospecific oxidation of primary alcohols to their corresponding aldehydes. The protein contains an unusual covalent thioether bond between a tyrosine, which acts as a radical center during the two-electron reaction, and a cysteine. The enzyme is produced in a precursor form lacking the thioether bond and also possessing an additional 17-aa pro-sequence at the N terminus. Previous work has shown that the aerobic addition of Cu(2+) to the precursor is sufficient to generate fully processed mature enzyme. The structure of the precursor protein has been determined to 1.4 A, revealing the location of the pro-sequence and identifying structural differences between the precursor and the mature protein. Structural alignment of the precursor and mature forms of galactose oxidase shows that five regions of main chain and some key residues of the active site differ significantly between the two forms. The precursor structure provides a starting point for modeling the chemistry of thioether bond formation and pro-sequence cleavage.
- Astbury Centre for Structural Molecular Biology, School of Biochemistry and Molecular Biology, University of Leeds, Leeds, LS2 9JT, United Kingdom.
Organizational Affiliation: