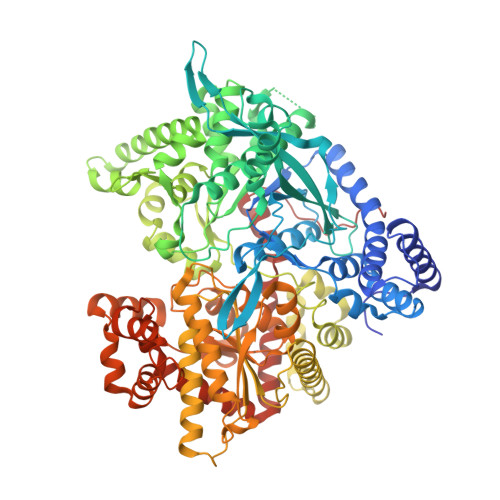
Binding of N-acetyl-N '-beta-D-glucopyranosyl urea and N-benzoyl-N '-beta-D-glucopyranosyl urea to glycogen phosphorylase b: kinetic and crystallographic studies.
Oikonomakos, N.G., Kosmopoulou, M., Zographos, S.E., Leonidas, D.D., Chrysina, E.D., Somsak, L., Nagy, V., Praly, J.P., Docsa, T., Toth, B., Gergely, P.(2002) Eur J Biochem 269: 1684-1696
- PubMed: 11895439 Search on PubMed
- DOI: https://doi.org/10.1046/j.1432-1327.2002.02813.x
- Primary Citation Related Structures:
1K06, 1K08, 1KTI - PubMed Abstract:
Two substituted ureas of beta-D-glucose, N-acetyl-N'-beta-D-glucopyranosyl urea (Acurea) and N-benzoyl-N'-beta-D-glucopyranosyl urea (Bzurea), have been identified as inhibitors of glycogen phosphorylase, a potential target for therapeutic intervention in type 2 diabetes. To elucidate the structural basis of inhibition, we determined the structure of muscle glycogen phosphorylase b (GPb) complexed with the two compounds at 2.0 A and 1.8 A resolution, respectively. The structure of the GPb-Acurea complex reveals that the inhibitor can be accommodated in the catalytic site of T-state GPb with very little change in the tertiary structure. The glucopyranose moiety makes the standard hydrogen bonds and van der Waals contacts as observed in the GPb-glucose complex, while the acetyl urea moiety is in a favourable electrostatic environment and makes additional polar contacts with the protein. The structure of the GPb-Bzurea complex shows that Bzurea binds tightly at the catalytic site and induces substantial conformational changes in the vicinity of the catalytic site. In particular, the loop of the polypeptide chain containing residues 282-287 shifts 1.3-3.7 A (Calpha atoms) to accommodate Bzurea. Bzurea can also occupy the new allosteric site, some 33 A from the catalytic site, which is currently the target for the design of antidiabetic drugs.
- Institute of Biological Research and Biotechnology, The National Hellenic Research Foundation, Athens, Greece. ngo@eie.gr
Organizational Affiliation: