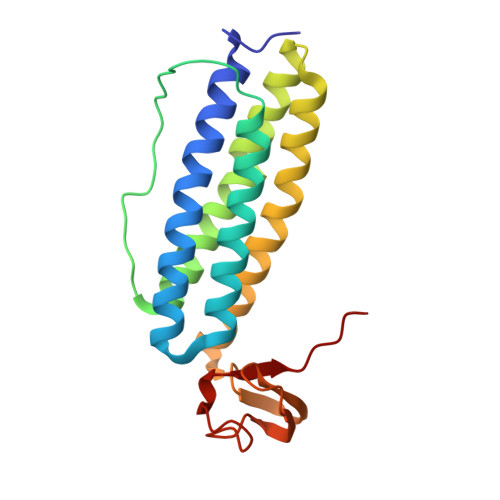
Crystal structure studies on rubrerythrin: enzymatic activity in relation to the zinc movement.
Li, M., Liu, M.Y., LeGall, J., Gui, L.L., Liao, J., Jiang, T., Zhang, J.P., Liang, D.C., Chang, W.R.(2003) J Biol Inorg Chem 8: 149-155
- PubMed: 12459910 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-002-0400-0
- Primary Citation Related Structures:
1JYB - PubMed Abstract:
Rubrerythrin (Rr) is a non-heme iron protein isolated from anaerobic sulfate-reducing bacteria. Rr is a dimeric molecule, each monomer contains a Fe(SCys)(4) center in the C-terminal domain and a binuclear metal center in the N-terminal domain. Rr structures with different protein sources and/or preparation procedures have been studied. Two Rr crystal structures have been solved with significant differences in their binuclear metal centers. The first structure, which was obtained from expressed protein under aerobic conditions, has a diiron-oxo center. The second structure, which was obtained from native protein of Desulfovibrio vulgaris under aerobic conditions, has an Fe-Zn center with the zinc position differing from the corresponding iron position in the former structure by approximately 2 A. The crystal structures of Rr isolated from D. vulgaris (Hildenborough, NCIB 8303), the same as the second structured but prepared under anaerobic conditions, are reported in this paper. The binuclear metal center in these structures is an Fe-Zn center. When the crystal was exposed to air, the zinc atom moved gradually, approximately 2 A, accompanied by the entrance of a water molecule (or hydroxyl group) and changes in the binuclear metal center microenvironment. This finding can explain the differences between the two different structures. The results suggest that the zinc movement may be related to the enzymatic activity of Rr.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Chaoyang District, Beijing 100101, China.
Organizational Affiliation: