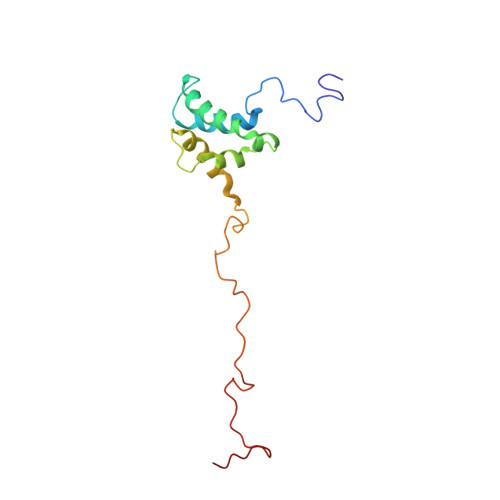
Three-dimensional structure of the HTLV-II matrix protein and comparative analysis of matrix proteins from the different classes of pathogenic human retroviruses.
Christensen, A.M., Massiah, M.A., Turner, B.G., Sundquist, W.I., Summers, M.F.(1996) J Mol Biology 264: 1117-1131
- PubMed: 9000634
- DOI: https://doi.org/10.1006/jmbi.1996.0700
- Primary Citation Related Structures:
1JVR - PubMed Abstract:
The matrix protein performs similar roles in all retroviruses, initially directing membrane localization of the assembling viral particle and subsequently forming a stable structural shell associated with the inner surface of the mature viral membrane. Although conserved structural elements are likely to perform these functions in all retroviral matrix proteins, invariant motifs are not evident at the primary sequence level and three-dimensional structures have been available for only the primate lentiviral matrix proteins. We have therefore used NMR spectroscopy to determine the structure of the matrix protein from human T-cell leukemia virus type II (HTLV-II), a member of the human oncovirus subclass of retroviruses. A total of 577 distance restraints were used to build 20 refined models that superimpose with an rmsd of 0.71 A for the backbone atoms of the structured regions. The globular HTLV-II matrix structure is composed of four alpha-helices and a 3(10) helix. Exposed basic residues near the C terminus of helix II form a putative membrane binding surface which could act in concert with the N-terminal myristoyl group to anchor the protein on the viral membrane surface. Clear structural similarities between the HTLV-II and HIV-1 matrix proteins suggest that the topology and exposed cationic membrane binding surface are likely to be conserved features of retroviral matrix proteins.
- Department of Biochemistry, University of Utah, Salt Lake City 84132, USA.
Organizational Affiliation: