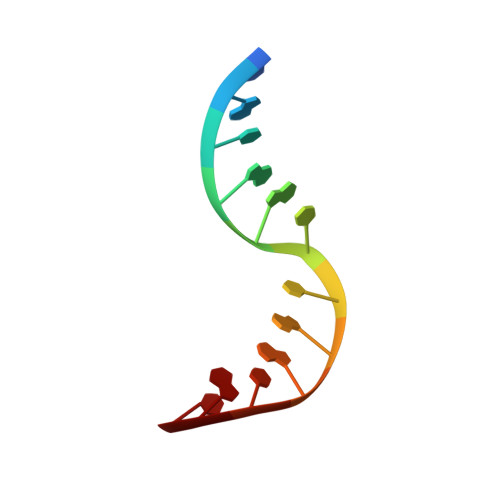
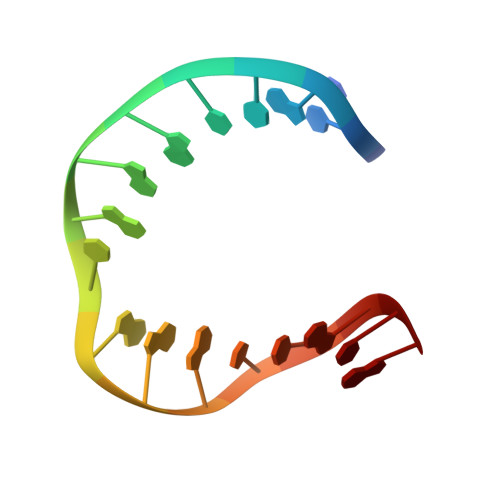
Solution structure of dAATAA and dAAUAA DNA bulges.
Gollmick, F.A., Lorenz, M., Dornberger, U., von Langen, J., Diekmann, S., Fritzsche, H.(2002) Nucleic Acids Res 30: 2669-2677
- PubMed: 12060684
- DOI: https://doi.org/10.1093/nar/gkf375
- Primary Citation Related Structures:
1JRV, 1JRW, 1JS5, 1JS7 - PubMed Abstract:
The NMR structure analysis is described for two DNA molecules of identical stem sequences with a five base loop containing a pyrimidine, thymin or uracil, in between purines. These five unpaired nucleotides are bulged out and are known to induce a kink in the duplex structure. The dAATAA bulge DNA is kinked between the third and the fourth nucleotide. This contrasts with the previously studied dAAAAA bulge DNA where we found a kink between the fourth and fifth nucleotide. The total kinking angle is approximately 104 degrees for the dAATAA bulge. The findings were supported by electrophoretic data and fluorescence resonance energy transfer measurements of a similar DNA molecule end-labeled by suitable fluorescent dyes. For the dAAUAA bulge the NMR data result in a similar structure as reported for the dAATAA bulge with a kinking angle of approximately 87 degrees. The results are discussed in comparison with a rAAUAA RNA bulge found in a group I intron. Generally, the sequence-dependent structure of bulges is important to understand the role of DNA bulges in protein recognition.
- Institut für Molekularbiologie, Friedrich-Schiller-Universität, Winzerlaer Strasse 10, D-07745 Jena, Germany.
Organizational Affiliation: