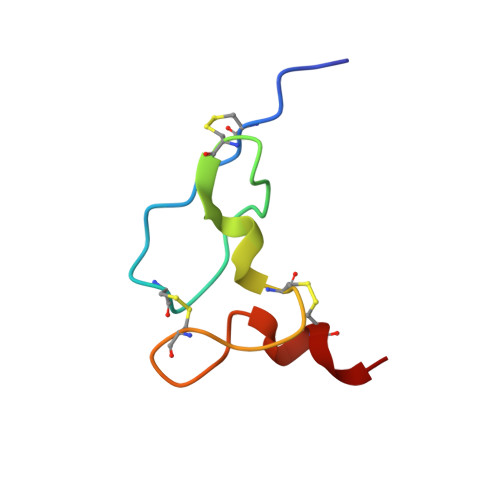
Solution structure of the viral receptor domain of Tva and its implications in viral entry.
Wang, Q.Y., Huang, W., Dolmer, K., Gettins, P.G., Rong, L.(2002) J Virol 76: 2848-2856
- PubMed: 11861852
- DOI: https://doi.org/10.1128/jvi.76.6.2848-2856.2002
- Primary Citation of Related Structures:
1JRF - PubMed Abstract:
Tva is the cellular receptor for subgroup A avian sarcoma and leukosis virus (ASLV-A). The viral receptor function of Tva is determined by a 40-residue, cysteine-rich motif called the LDL-A module. Here we report the solution structure of the LDL-A module of Tva, determined by nuclear magnetic resonance (NMR) spectroscopy. Although the carboxyl terminus of the Tva LDL-A module has a structure similar to those of other reported LDL-A modules, the amino terminus adopts a different conformation. The LDL-A module of Tva does not contain the signature antiparallel beta-sheet observed in other LDL-A modules, and it is more flexible than other reported LDL-A modules. The LDL-A structure of Tva provides mechanistic insights into how a simple viral receptor functions in retrovirus entry. The side chains of H38 and W48 of Tva, which have been identified as viral contact residues by mutational analysis, are solvent exposed, suggesting that they are directly involved in EnvA binding. However, the side chain of L34, another potential viral contact residue identified previously, is buried inside of the module and forms the hydrophobic core with other residues. Thus L34 likely stabilizes the Tva structure but is not a viral interaction determinant. In addition, we propose that the flexible amino-terminal region of Tva plays an important role in determining specificity in the Tva-EnvA interaction.
- Department of Microbiology and Immunology, College of Medicine, University of Illinois at Chicago, Chicago, Illinois 60612, USA.
Organizational Affiliation: