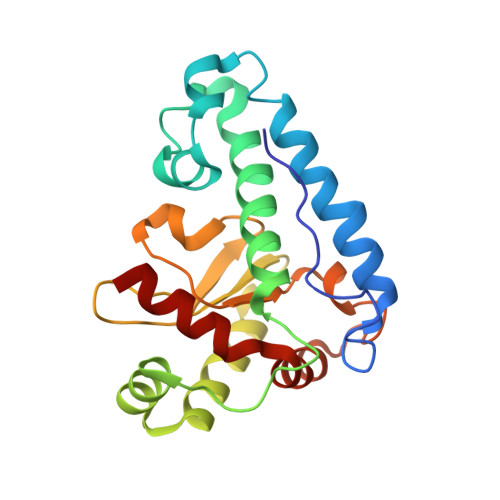
Three-dimensional structure of manganese superoxide dismutase from Bacillus halodenitrificans, a component of the so-called "green protein".
Liao, J., Liu, M.Y., Chang, T., Li, M., Le Gall, J., Gui, L.L., Zhang, J.P., Jiang, T., Liang, D.C., Chang, W.R.(2002) J Struct Biol 139: 171-180
- PubMed: 12457847 Search on PubMed
- DOI: https://doi.org/10.1016/s1047-8477(02)00531-2
- Primary Citation Related Structures:
1JR9 - PubMed Abstract:
A so-called "green protein" has been purified from a moderate halophilic eubacterium, Bacillus halodenitrificans (ATCC 49067), under anaerobic conditions. The protein, which might play an important role in denitrification, dissociates mainly into two components after exposure to air: a manganese superoxide dismutase (GP-MnSOD) and a nucleoside diphosphate kinase. As a first step in elucidating the overall structure of the green protein and the role of each component, the 2.8-A resolution crystal structure of GP-MnSOD was determined. Compared with other manganese dismutases, GP-MnSOD shows two significant characteristics. The first is that the entrance to its substrate channel has an additional basic residue-Lys38. The second is that its surface is decorated with an excess of acidic over basic residues. All these structural features may be related to GP-MnSOD's high catalytic activity and its endurance against the special cytoplasm of B. halodenitrificans. The structure of GP-MnSOD provides the basis for recognizing its possible role and assembly state in the green protein.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, 15 Datun Road, Chaoyang District, Beijing, China.
Organizational Affiliation: