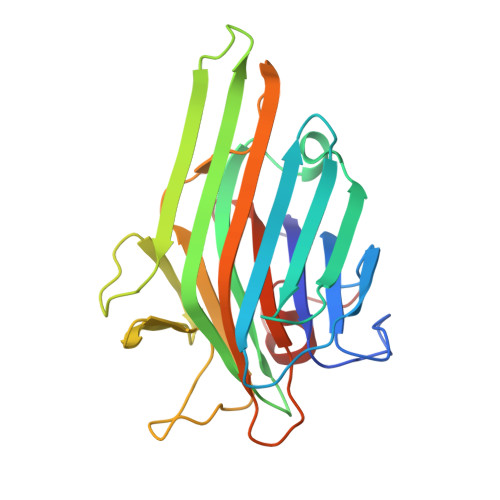
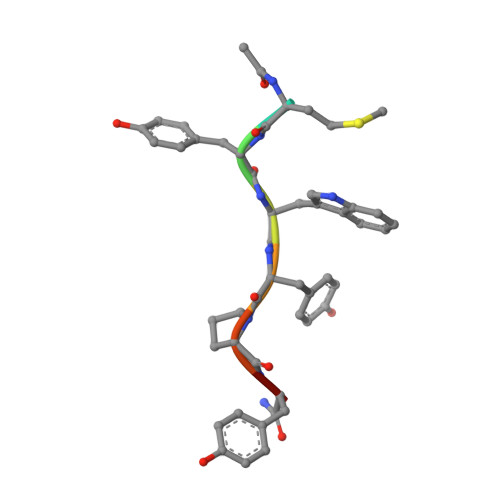
Enhanced binding of a rationally designed peptide ligand of concanavalin a arises from improved geometrical complementarity.
Jain, D., Kaur, K.J., Salunke, D.M.(2001) Biochemistry 40: 12059-12066
- PubMed: 11580281 Search on PubMed
- DOI: https://doi.org/10.1021/bi011254f
- Primary Citation Related Structures:
1JOJ - PubMed Abstract:
The structural basis of affinity enhancement was addressed by analyzing the interactions between concanavalin A and the carbohydrate-mimicking peptide ligands. Based on the crystal structures of concanavalin A in complex with these peptides [Jain, D., Kaur, K. J., Sundaravadivel, B., and Salunke, D. M. (2000) J. Biol. Chem. 275, 16098-16102; Jain, D., Kaur, K. J., and Salunke, D. M. (2001) Biophys. J. 80, 2912-2921], a high-affinity analogue was designed. This analogue (acetyl-MYWYPY-amide) binds to the lectin with 32-fold enhanced affinity compared to the corresponding precursor peptides. The crystal structure of concanavalin A bound to the designed peptide has been determined. A peptide molecule binds to each of the crystallographically independent monomers of the tetrameric lectin. The four bound peptide molecules exhibit two major conformations both of which are extended. Unlike in the case of other concanavalin A binding peptides, the structural variations within different conformers of this analogue are marginal. It is apparent that the deletion of the structurally variable region of the larger peptides has led to an improved complementarity and increased buried surface area in the case of the designed peptide. The crystal structure also showed the formation of two backbone hydrogen bonds between the ligand and the ligate which were not present in the complexes of the precursor peptides. The observed structural features adequately explain the enhanced binding of the designed analogue.
- Structural Biology Unit, National Institute of Immunology, Aruna Asaf Ali Marg, New Delhi 110 067, India.
Organizational Affiliation: