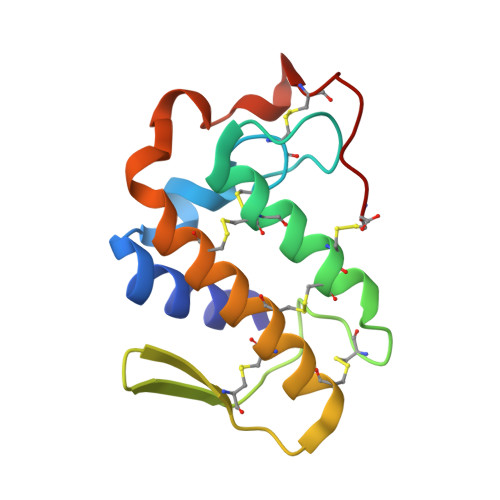
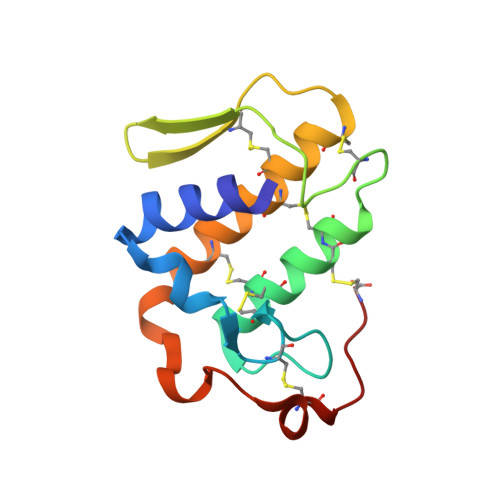
Structure of the neurotoxic complex vipoxin at 1.4 A resolution.
Banumathi, S., Rajashankar, K.R., Notzel, C., Aleksiev, B., Singh, T.P., Genov, N., Betzel, C.(2001) Acta Crystallogr D Biol Crystallogr 57: 1552-1559
- PubMed: 11679719
- DOI: https://doi.org/10.1107/s0907444901013543
- Primary Citation of Related Structures:
1JLT - PubMed Abstract:
Vipoxin is a neurotoxic postsynaptic heterodimeric complex from the venom of Vipera ammodytes meridionalis, the most toxic snake in Europe. It consists of a basic and highly toxic phospholipase A(2) and an acidic non-toxic protein inhibitor. The two polypeptide chains have the same chain length and share 62% amino-acid identity. Vipoxin is a unique example of evolution of the catalytic and toxic phospholipase A(2) functions into inhibitory and non-toxic functions. The crystal structure of the complex has been determined by the molecular-replacement method and refined to 1.4 A resolution to an R factor of 18.2%. The complex formation decreases the accessible surface area of the two subunits by approximately 1480 A(2), which results in a reduction of toxicity and catalytic activity. The catalytic and substrate-binding sites of the vipoxin phospholipase A(2) are identical or similar to those of other group I/II enzymes. Two 2-methyl-2,4-pentanediol molecules are present in the hydrophobic channel close to the active site. The two subunits lack calcium ions. The negatively charged Asp49 of the phospholipase A(2), which participates in the Ca(2+)-binding sites of other snake-venom phospholipase A(2)s, is neutralized by the side chain of Lys69 from the inhibitor. Attempts have been made to identify the toxicity region and to explain the reduced catalytic activity and toxicity of the phospholipase A(2) subunit.
- Institute of Medical Biochemistry and Molecular Biology, c/o DESY, Building 22a, Notkestrasse 85, 22603 Hamburg, Germany.
Organizational Affiliation: