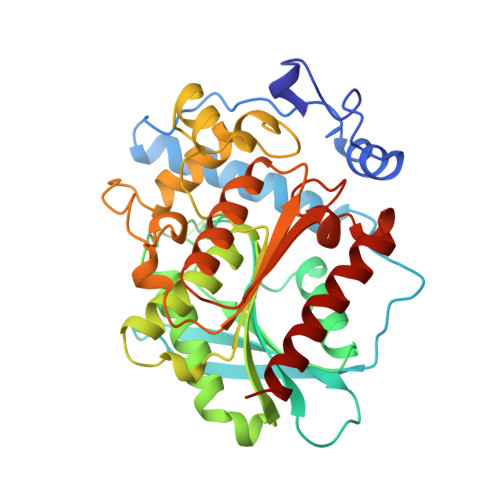
Crystal structure of brefeldin A esterase, a bacterial homolog of the mammalian hormone-sensitive lipase.
Wei, Y., Contreras, J.A., Sheffield, P., Osterlund, T., Derewenda, U., Kneusel, R.E., Matern, U., Holm, C., Derewenda, Z.S.(1999) Nat Struct Biol 6: 340-345
- PubMed: 10201402 Search on PubMed
- DOI: https://doi.org/10.1038/7576
- Primary Citation Related Structures:
1JKM - PubMed Abstract:
Brefeldin A esterase (BFAE), a detoxifying enzyme isolated from Bacillus subtilis, hydrolyzes and inactivates BFA, a potent fungal inhibitor of intracellular vesicle-dependent secretory transport and poliovirus RNA replication. We have solved the crystal structure of BFAE and we discovered that the previously reported amino acid sequence was in serious error due to frame shifts in the cDNA sequence. The correct sequence, inferred from the experimentally phased electron density map, revealed that BFAE is a homolog of the mammalian hormone sensitive lipase (HSL). It is a canonical alpha/beta hydrolase with two insertions forming the substrate binding pocket. The enzyme contains a lipase-like catalytic triad, Ser 202, Asp 308 and His 338, consistent with mutational studies that implicate the homologous Ser 424, Asp 693 and His 723 in the catalytic triad in human HSL.
- Department of Molecular Physiology and Biological Physics, University of Virginia, Charlottesville 22906-0011, USA.
Organizational Affiliation: