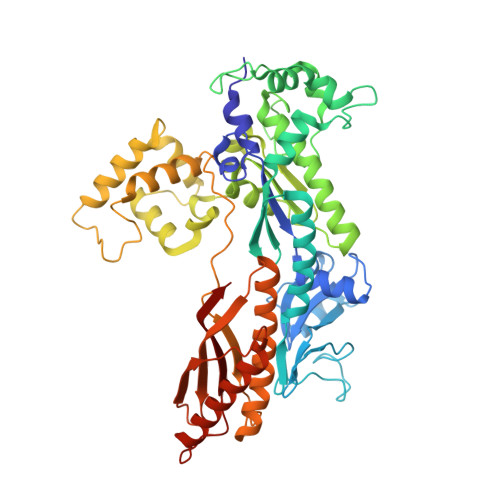
Structure of the catalytic core of S. cerevisiae DNA polymerase eta: implications for translesion DNA synthesis
Trincao, J., Johnson, R.E., Escalante, C.R., Prakash, S., Prakash, L., Aggarwal, A.K.(2001) Mol Cell 8: 417-426
- PubMed: 11545743 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(01)00306-9
- Primary Citation Related Structures:
1JIH - PubMed Abstract:
DNA polymerase eta is unique among eukaryotic polymerases in its proficient ability to replicate through a variety of distorting DNA lesions. We report here the crystal structure of the catalytic core of S. cerevisiae DNA polymerase eta, determined at 2.25A resolution. The structure reveals a novel polydactyl right hand-shaped molecule with a unique polymerase-associated domain. We identify the catalytic residues and show that the fingers and thumb domains are unusually small and stubby. In particular, the unexpected absence of helices "O" and "O1" in the fingers domain suggests that openness of the active site is the critical feature which enables DNA polymerase eta to replicate through DNA lesions such as a UV-induced cis-syn thymine-thymine dimer.
- Structural Biology Program, Department of Physiology and Biophysics, Mount Sinai School of Medicine, New York, NY 10029, USA.
Organizational Affiliation: