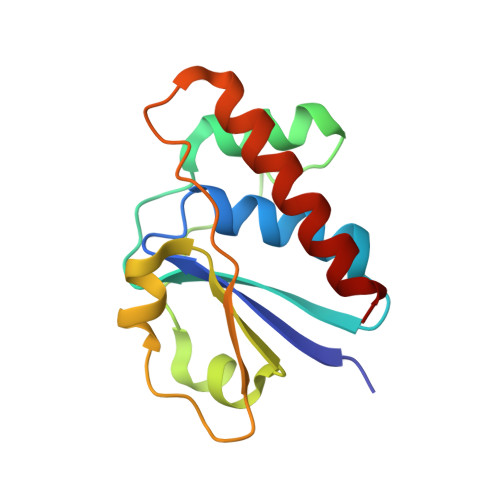
Arsenate reductase from S. aureus plasmid pI258 is a phosphatase drafted for redox duty.
Zegers, I., Martins, J.C., Willem, R., Wyns, L., Messens, J.(2001) Nat Struct Biol 8: 843-847
- PubMed: 11573087 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1001-843
- Primary Citation Related Structures:
1JF8, 1JFV - PubMed Abstract:
Arsenate reductase (ArsC) from Staphylococcus aureus plasmid pI258 plays a role in bacterial heavy metal resistance and catalyzes the reduction of arsenate to arsenite. The structures of the oxidized and reduced forms of ArsC were solved. ArsC has the PTPase I fold typical for low molecular weight tyrosine phosphatases (LMW PTPases). Remarkably, kinetic experiments show that pI258 ArsC also catalyzes the tyrosine phosphatase reaction in addition to arsenate reduction. These results provide evidence that ArsC from pI258 evolved from LMW PTPase by the grafting of a redox function onto a pre-existing catalytic site and that its evolutionary origin is different from those of arsenate reductases from Escherichia coli plasmid R773 and from Saccharomyces cerevisiae. The mechanism proposed here for the catalysis of arsenate reduction by pI258 ArsC involves a nucleophilic attack by Cys 10 on arsenate, the formation of a covalent intermediate and the transport of oxidative equivalents by a disulfide cascade. The reaction is associated with major structural changes in the ArsC.
- Dienst Ultrastruktuur, Vlaams interuniversitair Instituut voor Biotechnologie (VIB), Vrije Universiteit Brussel, Paardenstraat 65, B-1640 St. Genesius Rode, Belgium. igzegers@vub.ac.be
Organizational Affiliation: