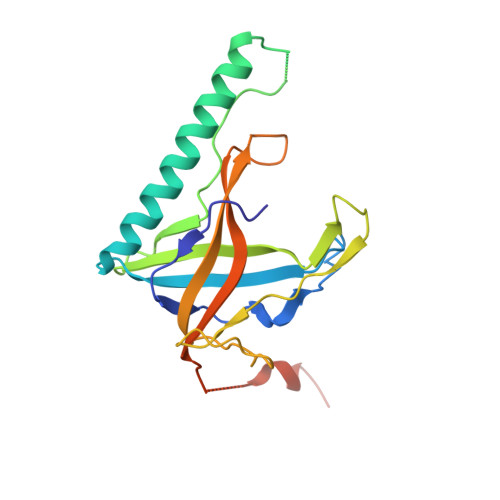
Structure of the gene 2.5 protein, a single-stranded DNA binding protein encoded by bacteriophage T7.
Hollis, T., Stattel, J.M., Walther, D.S., Richardson, C.C., Ellenberger, T.(2001) Proc Natl Acad Sci U S A 98: 9557-9562
- PubMed: 11481454 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.171317698
- Primary Citation Related Structures:
1JE5 - PubMed Abstract:
The gene 2.5 protein (gp2.5) of bacteriophage T7 is a single-stranded DNA (ssDNA) binding protein that has essential roles in DNA replication and recombination. In addition to binding DNA, gp2.5 physically interacts with T7 DNA polymerase and T7 primase-helicase during replication to coordinate events at the replication fork. We have determined a 1.9-A crystal structure of gp2.5 and show that it has a conserved OB-fold (oligosaccharide/oligonucleotide binding fold) that is well adapted for interactions with ssDNA. Superposition of the OB-folds of gp2.5 and other ssDNA binding proteins reveals a conserved patch of aromatic residues that stack against the bases of ssDNA in the other crystal structures, suggesting that gp2.5 binds to ssDNA in a similar manner. An acidic C-terminal extension of the gp2.5 protein, which is required for dimer formation and for interactions with the T7 DNA polymerase and the primase-helicase, appears to be flexible and may act as a switch that modulates the DNA binding affinity of gp2.5.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115, USA.
Organizational Affiliation: