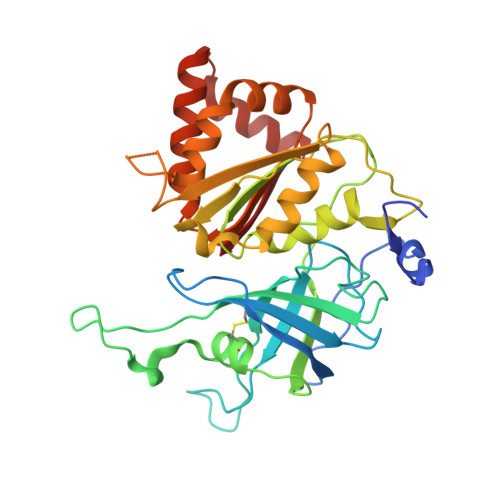
Biochemical and crystallographic characterization of ferredoxin-NADP(+) reductase from nonphotosynthetic tissues.
Aliverti, A., Faber, R., Finnerty, C.M., Ferioli, C., Pandini, V., Negri, A., Karplus, P.A., Zanetti, G.(2001) Biochemistry 40: 14501-14508
- PubMed: 11724563
- DOI: https://doi.org/10.1021/bi011224c
- Primary Citation of Related Structures:
1JB9, 3LVB - PubMed Abstract:
Distinct forms of ferredoxin-NADP(+) reductase are expressed in photosynthetic and nonphotosynthetic plant tissues. Both enzymes catalyze electron transfer between NADP(H) and ferredoxin; whereas in leaves the enzyme transfers reducing equivalents from photoreduced ferredoxin to NADP(+) in photosynthesis, in roots it has the opposite physiological role, reducing ferredoxin at the expense of NADPH mainly for use in nitrate assimilation. Here, structural and kinetic properties of a nonphotosynthetic isoform were analyzed to define characteristics that may be related to tissue-specific function. Compared with spinach leaf ferredoxin-NADP(+) reductase, the recombinant corn root isoform showed a slightly altered absorption spectrum, a higher pI, a >30-fold higher affinity for NADP(+), greater susceptibility to limited proteolysis, and an approximately 20 mV more positive redox potential. The 1.7 A resolution crystal structure is very similar to the structures of ferredoxin-NADP(+) reductases from photosynthetic tissues. Four distinct structural features of this root ferredoxin-NADP(+) reductases are an alternate conformation of the bound FAD molecule, an alternate path for the amino-terminal extension, a disulfide bond in the FAD-binding domain, and changes in the surface that binds ferredoxin.
- Dipartimento di Fisiologia e Biochimica Generali, Università degli Studi di Milano, Via Celoria 26, 20133 Milano, Italy.
Organizational Affiliation: