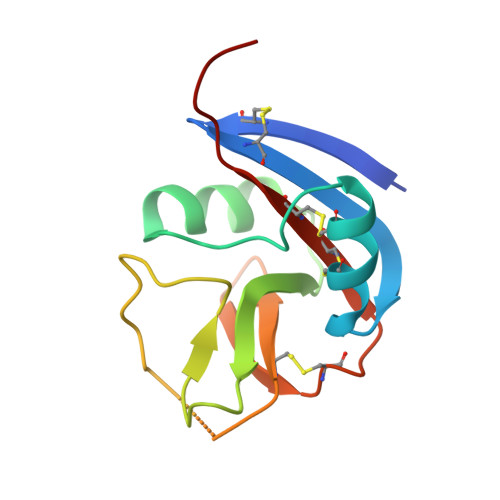
Crystal structure of the Ly49I natural killer cell receptor reveals variability in dimerization mode within the Ly49 family.
Dimasi, N., Sawicki, M.W., Reineck, L.A., Li, Y., Natarajan, K., Margulies, D.H., Mariuzza, R.A.(2002) J Mol Biology 320: 573-585
- PubMed: 12096910
- DOI: https://doi.org/10.1016/s0022-2836(02)00498-9
- Primary Citation Related Structures:
1JA3 - PubMed Abstract:
Natural killer (NK) cells play a crucial role in the detection and destruction of virally infected and tumor cells during innate immune responses. The cytolytic activity of NK cells is regulated through a balance of inhibitory and stimulatory signals delivered by NK receptors that recognize classical major histocompatabilty complex class I (MHC-I) molecules, or MHC-I homologs such as MICA, on target cells. The Ly49 family of NK receptors (Ly49A through W), which includes both inhibitory and activating receptors, are homodimeric type II transmembrane glycoproteins, with each subunit composed of a C-type lectin-like domain tethered to the membrane by a stalk region. We have determined the crystal structure, at 3.0 A resolution, of the murine inhibitory NK receptor Ly49I. The Ly49I monomer adopts a fold similar to that of other C-type lectin-like NK receptors, including Ly49A, NKG2D and CD69. However, the Ly49I monomers associate in a manner distinct from that of these other NK receptors, forming a more open dimer. As a result, the putative MHC-binding surfaces of the Ly49I dimer are spatially more distant than the corresponding surfaces of Ly49A or NKG2D. These structural differences probably reflect the fundamentally different ways in which Ly49 and NKG2D receptors recognize their respective ligands: whereas the single MICA binding site of NKG2D is formed by the precise juxtaposition of two monomers, each Ly49 monomer contains an independent binding site for MHC-I. Hence, the structural constraints on dimerization geometry may be relatively relaxed within the Ly49 family. Such variability may enable certain Ly49 receptors, like Ly49I, to bind MHC-I molecules bivalently, thereby stabilizing receptor-ligand interactions and enhancing signal transmission to the NK cell.
- W.M. Keck Laboratory for Structural Biology, Center for Advanced Research in Biotechnology, University of Maryland Biotechnology Institute, 9600 Gudelsky Drive, Rockville, MD 20850, USA.
Organizational Affiliation: