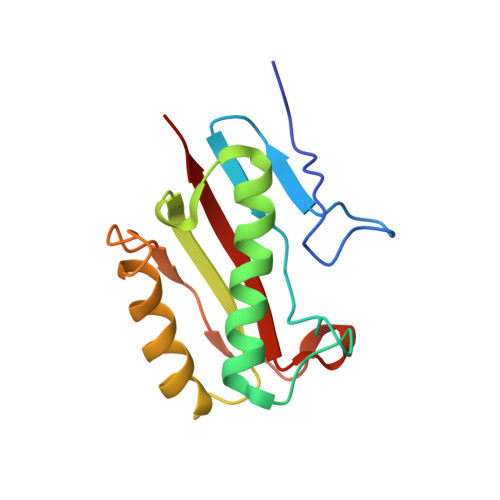
Solution Structure and Functional Ligand Screening of HI0719, a Highly Conserved Protein from Bacteria to Humans in the YjgF/YER057c/UK114 Family
Parsons, L., Bonander, N., Eisenstein, E., Gilson, M., Kairys, V., Orban, J.(2003) Biochemistry 42: 80-89
- PubMed: 12515541
- DOI: https://doi.org/10.1021/bi020541w
- Primary Citation of Related Structures:
1J7H - PubMed Abstract:
HI0719 belongs to a large family of highly conserved proteins with no definitive molecular function and is found in organisms ranging from bacteria to humans. We describe the NMR structure of HI0719, the first solution structure for a member of this family. The overall fold is similar to the crystal structures of two homologues, YabJ from Bacillus subtilis and YjgF from Escherichia coli, and all three structures are similar to that of chorismate mutase, although there is little sequence homology and no apparent functional connection. HI0719 is a homotrimer with a distinct cavity located at the subunit interface. Six of the seven invariant residues in the high identity group of proteins are located in this cavity, suggesting that this may be a binding site for small molecules. Using previously published observations about the biological role of HI0719 family members as a guide, over 100 naturally occurring small molecules or structural analogues were screened for ligand binding using NMR spectroscopy. The targeted screening approach identified six compounds that bind to HI0719 at the putative active site. Five of these compounds are either alpha-keto acids or alpha,beta-unsaturated acids, while the sixth compound is structurally similar. Previous studies have proposed that some HI0719 homologues may act on small molecules in the isoleucine biosynthetic path and, if this is correct, the ligand screening results presented here suggest that the interaction most likely occurs with 2-ketobutyrate and/or its unstable enamine precursor.
- Center for Advanced Research in Biotechnology, University of Maryland Biotechnology Institute, 9600 Gudelsky Drive, Rockville, Maryland 20850, USA.
Organizational Affiliation: