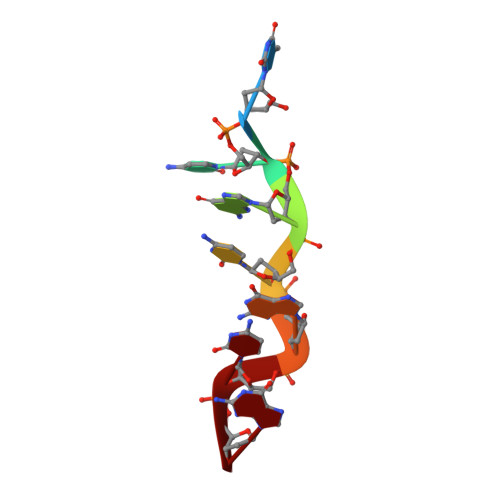
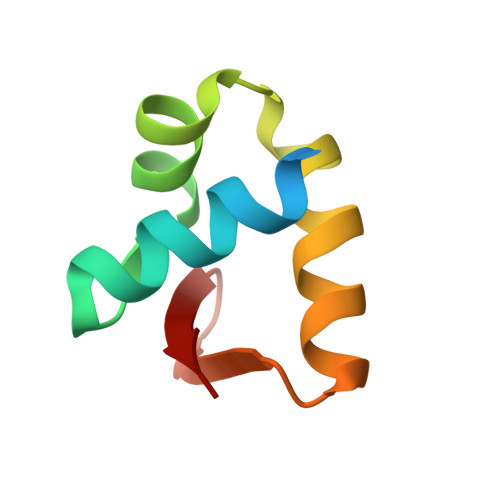
Structure of the DLM-1-Z-DNA complex reveals a conserved family of Z-DNA-binding proteins.
Schwartz, T., Behlke, J., Lowenhaupt, K., Heinemann, U., Rich, A.(2001) Nat Struct Biol 8: 761-765
- PubMed: 11524677 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0901-761
- Primary Citation Related Structures:
1J75 - PubMed Abstract:
The first crystal structure of a protein, the Z alpha high affinity binding domain of the RNA editing enzyme ADAR1, bound to left-handed Z-DNA was recently described. The essential set of residues determined from this structure to be critical for Z-DNA recognition was used to search the database for other proteins with the potential for Z-DNA binding. We found that the tumor-associated protein DLM-1 contains a domain with remarkable sequence similarities to Z alpha(ADAR). Here we report the crystal structure of this DLM-1 domain bound to left-handed Z-DNA at 1.85 A resolution. Comparison of Z-DNA binding by DLM-1 and ADAR1 reveals a common structure-specific recognition core within the binding domain. However, the domains differ in certain residues peripheral to the protein-DNA interface. These structures reveal a general mechanism of Z-DNA recognition, suggesting the existence of a family of winged-helix proteins sharing a common Z-DNA binding motif.
- Max-Delbrück-Centrum für Molekulare Medizin, Robert-Rössle-Str. 10, D-13125 Berlin, Germany. Thomas.Schwartz@mail.rockefeller.edu
Organizational Affiliation: