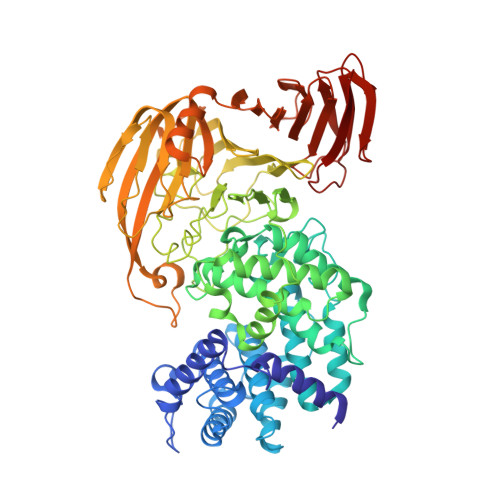
Crystal Structure of Bacillus sp. GL1 Xanthan Lyase, Which Acts on the Side Chains of Xanthan.
Hashimoto, W., Nankai, H., Mikami, B., Murata, K.(2003) J Biological Chem 278: 7663-7673
- PubMed: 12475987
- DOI: https://doi.org/10.1074/jbc.M208100200
- Primary Citation Related Structures:
1J0M, 1J0N - PubMed Abstract:
Xanthan lyase, a member of polysaccharide lyase family 8, is a key enzyme for complete depolymerization of a bacterial heteropolysaccharide, xanthan, in Bacillus sp. GL1. The enzyme acts exolytically on the side chains of the polysaccharide. The x-ray crystallographic structure of xanthan lyase was determined by the multiple isomorphous replacement method. The crystal structures of xanthan lyase and its complex with the product (pyruvylated mannose) were refined at 2.3 and 2.4 A resolution with final R-factors of 17.5 and 16.9%, respectively. The refined structure of the product-free enzyme comprises 752 amino acid residues, 248 water molecules, and one calcium ion. The enzyme consists of N-terminal alpha-helical and C-terminal beta-sheet domains, which constitute incomplete alpha(5)/alpha(5)-barrel and anti-parallel beta-sheet structures, respectively. A deep cleft is located in the N-terminal alpha-helical domain facing the interface between the two domains. Although the overall structure of the enzyme is basically the same as that of the family 8 lyases for hyaluronate and chondroitin AC, significant differences were observed in the loop structure over the cleft. The crystal structure of the xanthan lyase complexed with pyruvylated mannose indicates that the sugar-binding site is located in the deep cleft, where aromatic and positively charged amino acid residues are involved in the binding. The Arg(313) and Tyr(315) residues in the loop from the N-terminal domain and the Arg(612) residue in the loop from the C-terminal domain directly bind to the pyruvate moiety of the product through the formation of hydrogen bonds, thus determining the substrate specificity of the enzyme.
- Department of Basic and Applied Molecular Biotechnology, Division of Food and Biological Science, Graduate School of Agriculture, Kyoto, Uji, Kyoto 611-0011, Japan.
Organizational Affiliation: