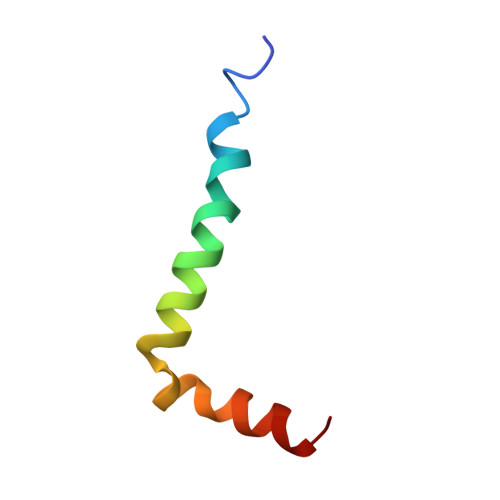
Solution structure of the Alzheimer amyloid beta-peptide (1-42) in an apolar microenvironment. Similarity with a virus fusion domain.
Crescenzi, O., Tomaselli, S., Guerrini, R., Salvadori, S., D'Ursi, A.M., Temussi, P.A., Picone, D.(2002) Eur J Biochem 269: 5642-5648
- PubMed: 12423364 Search on PubMed
- DOI: https://doi.org/10.1046/j.1432-1033.2002.03271.x
- Primary Citation Related Structures:
1IYT - PubMed Abstract:
The major components of neuritic plaques found in Alzheimer disease (AD) are peptides known as amyloid beta-peptides (Abeta), which derive from the proteolitic cleavage of the amyloid precursor proteins. In vitro Abeta may undergo a conformational transition from a soluble form to aggregated, fibrillary beta-sheet structures, which seem to be neurotoxic. Alternatively, it has been suggested that an alpha-helical form can be involved in a process of membrane poration, which would then trigger cellular death. Conformational studies on these peptides in aqueous solution are complicated by their tendency to aggregate, and only recently NMR structures of Abeta-(1-40) and Abeta-(1-42) have been determined in aqueous trifluoroethanol or in SDS micelles. All these studies hint to the presence of two helical regions, connected through a flexible kink, but it proved difficult to determine the length and position of the helical stretches with accuracy and, most of all, to ascertain whether the kink region has a preferred conformation. In the search for a medium which could allow a more accurate structure determination, we performed an exhaustive solvent scan that showed a high propensity of Abeta-(1-42) to adopt helical conformations in aqueous solutions of fluorinated alcohols. The 3D NMR structure of Abeta-(1-42) shows two helical regions encompassing residues 8-25 and 28-38, connected by a regular type I beta-turn. The surprising similarity of this structure, as well as the sequence of the C-terminal moiety, with those of the fusion domain of influenza hemagglutinin suggests a direct mechanism of neurotoxicity.
- Dipartimento di Chimica, Università degli Studi di Napoli 'Federico II', Italy.
Organizational Affiliation: