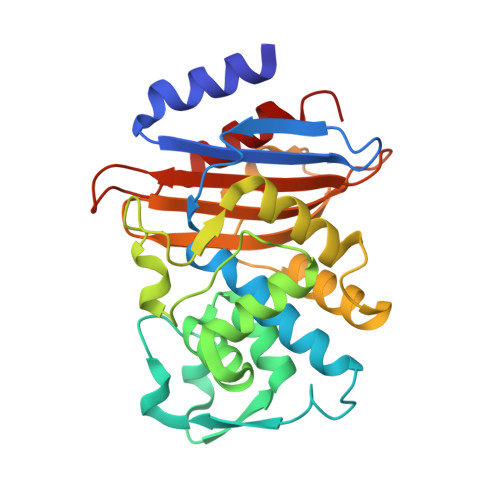
Acyl-intermediate Structures of the Extended-spectrum Class A beta -Lactamase, Toho-1, in Complex with Cefotaxime, Cephalothin, and Benzylpenicillin.
Shimamura, T., Ibuka, A., Fushinobu, S., Wakagi, T., Ishiguro, M., Ishii, Y., Matsuzawa, H.(2002) J Biological Chem 277: 46601-46608
- PubMed: 12221102 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M207884200
- Primary Citation Related Structures:
1IYO, 1IYP, 1IYQ - PubMed Abstract:
Bacterial resistance to beta-lactam antibiotics is a serious problem limiting current clinical therapy. The most common form of resistance is the production of beta-lactamases that inactivate beta-lactam antibiotics. Toho-1 is an extended-spectrum beta-lactamase that has acquired efficient activity not only to penicillins but also to cephalosporins including the expanded-spectrum cephalosporins that were developed to be stable in former beta-lactamases. We present the acyl-intermediate structures of Toho-1 in complex with cefotaxime (expanded-spectrum cephalosporin), cephalothin (non-expanded-spectrum cephalosporin), and benzylpenicillin at 1.8-, 2.0-, and 2.1-A resolutions, respectively. These structures reveal distinct features that can explain the ability of Toho-1 to hydrolyze expanded-spectrum cephalosporins. First, the Omega-loop of Toho-1 is displaced to avoid the steric contacts with the bulky side chain of cefotaxime. Second, the conserved residues Asn(104) and Asp(240) form unique interactions with the bulky side chain of cefotaxime to fix it tightly. Finally, the unique interaction between the conserved Ser(237) and cephalosporins probably helps to bring the beta-lactam carbonyl group to the suitable position in the oxyanion hole, thus increasing the cephalosporinase activity.
- Department of Biotechnology, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan.
Organizational Affiliation: