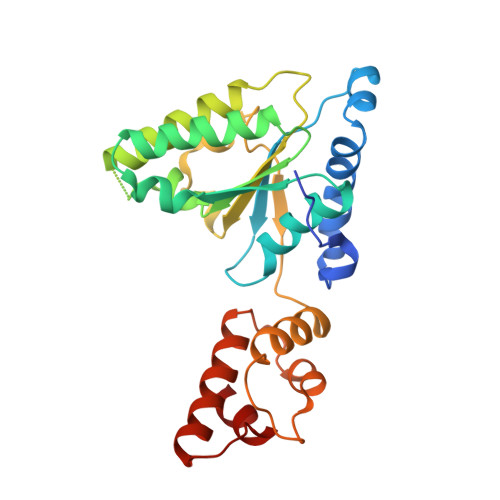
Hexameric ring structure of the ATPase domain of the membrane-integrated metalloprotease FtsH from Thermus thermophilus HB8
Niwa, H., Tsuchiya, D., Makyio, H., Yoshida, M., Morikawa, K.(2002) Structure 10: 1415-1423
- PubMed: 12377127 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00855-9
- Primary Citation Related Structures:
1IXZ, 1IY0, 1IY1, 1IY2 - PubMed Abstract:
FtsH is a cytoplasmic membrane-integrated, ATP-dependent metalloprotease, which processively degrades both cytoplasmic and membrane proteins in concert with unfolding. The FtsH protein is divided into the N-terminal transmembrane region and the larger C-terminal cytoplasmic region, which consists of an ATPase domain and a protease domain. We have determined the crystal structures of the Thermus thermophilus FtsH ATPase domain in the nucleotide-free and AMP-PNP- and ADP-bound states, in addition to the domain with the extra preceding segment. Combined with the mapping of the putative substrate binding region, these structures suggest that FtsH internally forms a hexameric ring structure, in which ATP binding could cause a conformational change to facilitate transport of substrates into the protease domain through the central pore.
- Department of Structural Biology, Biomolecular Engineering Research Institute, 6-2-3 Furuedai, Suita, Osaka, Japan.
Organizational Affiliation: