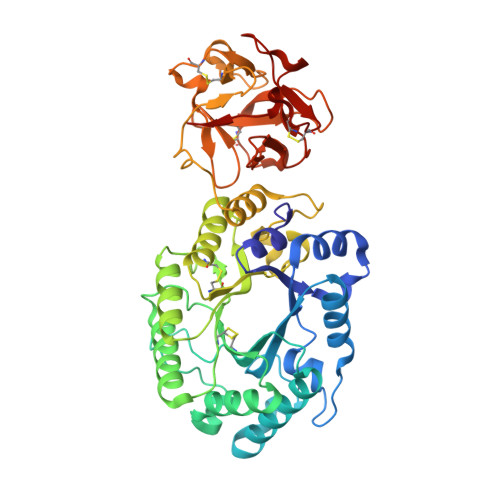
Crystal structures of the sugar complexes of Streptomyces olivaceoviridis E-86 xylanase: sugar binding structure of the family 13 carbohydrate binding module.
Fujimoto, Z., Kuno, A., Kaneko, S., Kobayashi, H., Kusakabe, I., Mizuno, H.(2002) J Mol Biology 316: 65-78
- PubMed: 11829503
- DOI: https://doi.org/10.1006/jmbi.2001.5338
- Primary Citation of Related Structures:
1ISV, 1ISW, 1ISX, 1ISY, 1ISZ, 1IT0 - PubMed Abstract:
The family 10 xylanase from Streptomyces olivaceoviridis E-86 contains a (beta/alpha)(8)-barrel as a catalytic domain, a family 13 carbohydrate binding module (CBM) as a xylan binding domain (XBD) and a Gly/Pro-rich linker between them. The crystal structure of this enzyme showed that XBD has three similar subdomains, as indicated by the presence of a triple-repeated sequence, forming a galactose binding lectin fold similar to that found in the ricin toxin B-chain. Comparison with the structure of ricin/lactose complex suggests three potential sugar binding sites in XBD. In order to understand how XBD binds to the xylan chain, we analyzed the sugar-complex structure by the soaking experiment method using the xylooligosaccharides and other sugars. In the catalytic cleft, bound sugars were observed in the xylobiose and xylotriose complex structures. In the XBD, bound sugars were identified in subdomains alpha and gamma in all of the complexes with xylose, xylobiose, xylotriose, glucose, galactose and lactose. XBD binds xylose or xylooligosaccharides at the same sugar binding sites as in the case of the ricin/lactose complex but its binding manner for xylose and xylooligosaccharides is different from the galactose binding mode in ricin, even though XBD binds galactose in the same manner as in the ricin/galactose complex. These different binding modes are utilized efficiently and differently to bind the long substrate to xylanase and ricin-type lectin. XBD can bind any xylose in the xylan backbone, whereas ricin-type lectin recognizes the terminal galactose to sandwich the large sugar chain, even though the two domains have the same family 13 CBM structure. Family 13 CBM has rather loose and broad sugar specificities and is used by some kinds of proteins to bind their target sugars. In such enzyme, XBD binds xylan, and the catalytic domain may assume a flexible position with respect to the XBD/xylan complex, inasmuch as the linker region is unstructured.
- Department of Biochemistry, National Institute of Agrobiological Sciences, Tsukuba, Ibaraki, 305-8602, Japan. zui@affrc.go.jp
Organizational Affiliation: