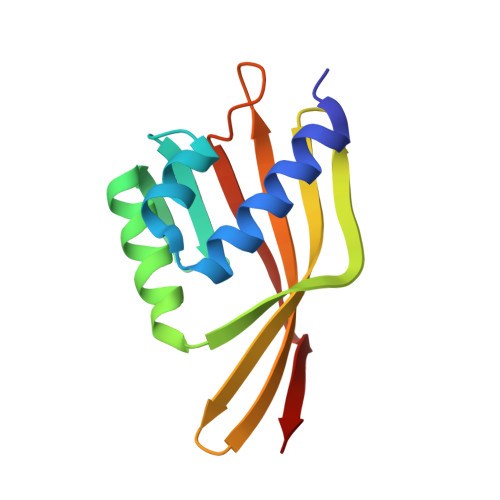
Solution structure of 3-oxo-delta5-steroid isomerase.
Wu, Z.R., Ebrahimian, S., Zawrotny, M.E., Thornburg, L.D., Perez-Alvarado, G.C., Brothers, P., Pollack, R.M., Summers, M.F.(1997) Science 276: 415-418
- PubMed: 9103200
- DOI: https://doi.org/10.1126/science.276.5311.415
- Primary Citation of Related Structures:
1ISK - PubMed Abstract:
The three-dimensional structure of the enzyme 3-oxo-delta5-steroid isomerase (E.C. 5.3.3.1), a 28-kilodalton symmetrical dimer, was solved by multidimensional heteronuclear magnetic resonance spectroscopy. The two independently folded monomers pack together by means of extensive hydrophobic and electrostatic interactions. Each monomer comprises three alpha helices and a six-strand mixed beta-pleated sheet arranged to form a deep hydrophobic cavity. Catalytically important residues Tyr14 (general acid) and Asp38 (general base) are located near the bottom of the cavity and positioned as expected from mechanistic hypotheses. An unexpected acid group (Asp99) is also located in the active site adjacent to Tyr14, and kinetic and binding studies of the Asp99 to Ala mutant demonstrate that Asp99 contributes to catalysis by stabilizing the intermediate.
- Howard Hughes Medical Institute and Department of Chemistry and Biochemistry, University of Maryland Baltimore County, 1000 Hilltop Circle, Baltimore, MD 21250.
Organizational Affiliation: