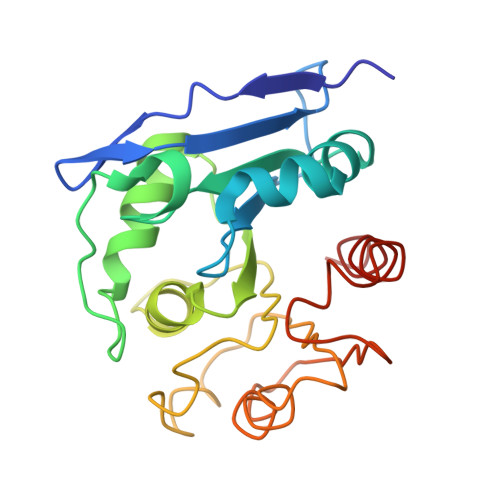
The crystal structure of CCG1/TAF(II)250-interacting factor B (CIB)
Padmanabhan, B., Kuzuhara, T., Adachi, N., Horikoshi, M.(2004) J Biological Chem 279: 9615-9624
- PubMed: 14672934 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M312165200
- Primary Citation Related Structures:
1IMJ - PubMed Abstract:
The general transcription initiation factor TFIID and its interactors play critical roles in regulating the transcription from both naked and chromatin DNA. We have isolated a novel TFIID interactor that we denoted as CCG1/TAF(II)250-interacting factor B (CIB). We show here that CIB activates transcription. To further understand the function of this protein, we determined its crystal structure at 2.2-Angstroms resolution. The tertiary structure of CIB reveals an alpha/beta-hydrolase fold that resembles structures in the prokaryotic alpha/beta-hydrolase family proteins. It is not similar in structure or primary sequence to any eukaryotic transcription or chromatin factors that have been reported to date. CIB possesses a conserved catalytic triad that is found in other alpha/beta-hydrolases, and our in vitro studies confirmed that it bears hydrolase activity. However, CIB differs from other alpha/beta-hydrolases in that it lacks a binding site excursion, which facilitates the substrate selectivity of the other alpha/beta-hydrolases. Further functional characterization of CIB based on its tertiary structure and through biochemical studies may provide novel insights into the mechanisms that regulate eukaryotic transcription.
- Horikoshi Gene Selector Project, Exploratory Research for Advanced Technology, Japan Science and Technology Corporation, 5-9-6 Tokodai, Tsukuba, Ibaraki 300-2635, Japan.
Organizational Affiliation: