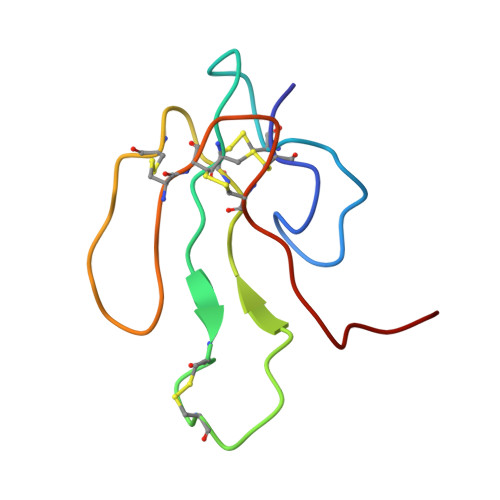
NMR structure of alpha-bungarotoxin free and bound to a mimotope of the nicotinic acetylcholine receptor.
Scarselli, M., Spiga, O., Ciutti, A., Bernini, A., Bracci, L., Lelli, B., Lozzi, L., Calamandrei, D., Di Maro, D., Klein, S., Niccolai, N.(2002) Biochemistry 41: 1457-1463
- PubMed: 11814338
- DOI: https://doi.org/10.1021/bi011012f
- Primary Citation of Related Structures:
1HOY, 1IK8, 1IKC, 1JBD - PubMed Abstract:
A combinatorial library approach was used to produce synthetic peptides mimicking the snake neurotoxin binding site of nicotinic receptors. Among the sequences, which inhibited binding of alpha-bungarotoxin to muscle and neuronal nicotinic receptors, HRYYESSLPWYPD, a 14-amino acid peptide with considerably higher toxin-binding affinity than the other synthesized peptides, was selected, and the structure of its complex with the toxin was analyzed by NMR. Comparison of the solution structure of the free toxin and its complex with this peptide indicated that complex formation induced extensive conformational rearrangements mainly at finger II and the carboxy terminus of the protein. The peptidyl residues P10 and Y4 seemed to be critical for peptide folding and complex stability, respectively. The latter residue of the peptide strongly interacted with the protein by entering a small pocket delimited by D30, C33, S34, R36, and V39 toxin side chains.
- Department of Molecular Biology and Biomolecular Structure Research Center, University of Siena, Italy.
Organizational Affiliation: