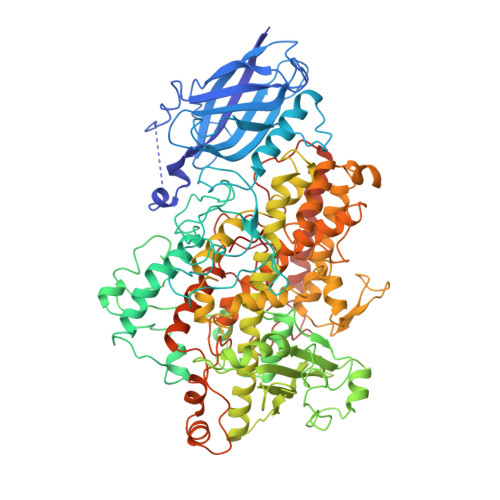
Three-dimensional structure of a purple lipoxygenase.
Skrzypczak-Jankun, E., Bross, R.A., Carroll, R.T., Dunham, W.R., Funk, M.O.(2001) J Am Chem Soc 123: 10814-10820
- PubMed: 11686682
- DOI: https://doi.org/10.1021/ja011759t
- Primary Citation Related Structures:
1IK3 - PubMed Abstract:
Polyunsaturated fatty acid metabolism is governed primarily by two enzymes, prostaglandin H synthase and lipoxygenase. The crystal structure of the metastable product-oxidized purple form of soybean lipoxygenase-3 was determined at 2.0 A resolution. The data reveal that the chromophore corresponds to an iron-peroxide complex, a potential intermediate in the catalyzed reaction. A significant alteration of the iron site accompanies the formation of the complex. The structure, the first for a fatty acid-lipoxygenase complex, also reveals an unexpected mode of binding, and identifies amino acid residues that may play significant roles in catalysis, regio- and stereoselectivity.
- Department of Chemistry, University of Toledo, 2801 West Bancroft Street, Toledo, OH 43606, USA. ejankun@protein.wo.utoledo.edu
Organizational Affiliation: