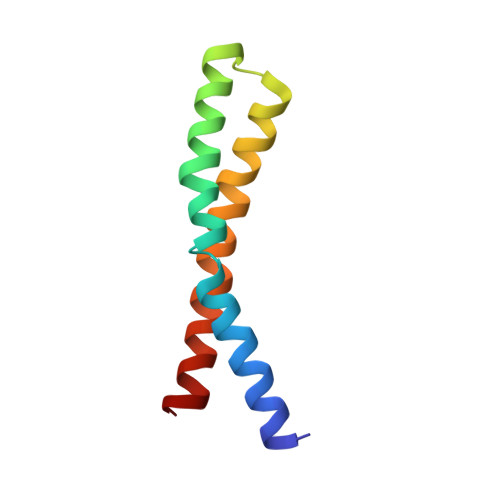
Structure of Ala(20) --> Pro/Pro(64) --> Ala substituted subunit c of Escherichia coli ATP synthase in which the essential proline is switched between transmembrane helices.
Dmitriev, O.Y., Fillingame, R.H.(2001) J Biological Chem 276: 27449-27454
- PubMed: 11331283 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M100762200
- Primary Citation Related Structures:
1IJP - PubMed Abstract:
The structure of the A20P/P64A mutated subunit c of Escherichia coli ATP synthase, in which the essential proline has been switched from residue 64 of the second transmembrane helix (TMH) to residue 20 of the first TMH, has been solved by (15)N,(1)H NMR in a monophasic chloroform/methanol/water (4:4:1) solvent mixture. The cA20P/P64A mutant grows as well as wild type, and the F(0)F(1) complex is fully functional in ATPase-coupled H(+) pumping. Residues 20 and 64 lie directly opposite to each other in the hairpin-like structure of wild type subunit c, and the prolinyl 64 residue is thought to induce a slight bend in TMH-2 such that it wraps around a more straightened TMH-1. In solution, the A20P/P64A substituted subunit c also forms a hairpin of two alpha-helices, with residues 41-45 forming a connecting loop as in the case of the wild type protein, but, in this case, Pro(20) induces a bend in TMH-1, which then packs against a more straightened TMH-2. The essential prolinyl residue, whether at position 64 or 20, lies close to the aspartyl 61 H(+) binding site. The prolinyl residue may introduce structural flexibility in this region of the protein, which may be necessary for the proposed movement of the alpha-helical segments during the course of the H(+) pumping catalytic cycle.
- Department of Biomolecular Chemistry, University of Wisconsin Medical School, Madison, Wisconsin 53706, USA.
Organizational Affiliation: