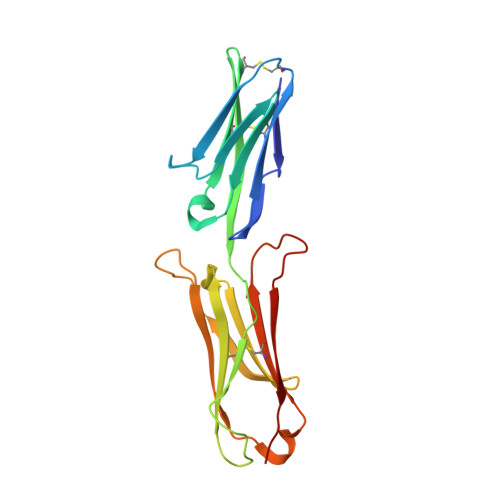
A new conformation of the integrin-binding fragment of human VCAM-1 crystallizes in a highly hydrated packing arrangement.
Taylor, P., Bilsland, M., Walkinshaw, M.D.(2001) Acta Crystallogr D Biol Crystallogr 57: 1579-1583
- PubMed: 11679722 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444901011209
- Primary Citation Related Structures:
1IJ9 - PubMed Abstract:
An X-ray crystal structure of two N-terminal integrin-binding IgSF domains of human VCAM-1 is reported. This new crystal form shows an unusual and highly hydrated packing arrangement in which over 80% of the crystal is occupied by solvent. The relative orientations of the two domains adopt a new intermediate conformation. The tilt angle between the two domains is 19.4 degrees, compared with other related structures that have tilt angles ranging from 7.3 to 39.9 degrees. An analysis of the torsion angles shows that residues Ile88, Tyr89, Ser90, Pro92 and Glu96 play a major role in defining the interdomain conformations.
- Structural Biochemistry Group, Institute of Cell and Molecular Biology, The University of Edinburgh, Michael Swann Building, King's Buildings, Edinburgh EH9 3JR, Scotland.
Organizational Affiliation: