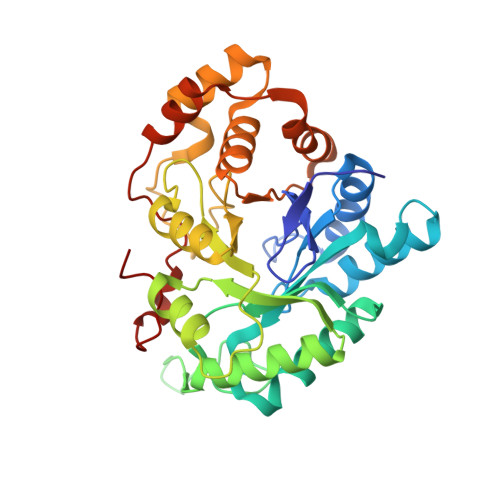
Crystal structure of human type III 3alpha-hydroxysteroid dehydrogenase/bile acid binding protein complexed with NADP(+) and ursodeoxycholate.
Jin, Y., Stayrook, S.E., Albert, R.H., Palackal, N.T., Penning, T.M., Lewis, M.(2001) Biochemistry 40: 10161-10168
- PubMed: 11513593 Search on PubMed
- DOI: https://doi.org/10.1021/bi010919a
- Primary Citation Related Structures:
1IHI - PubMed Abstract:
The crystal structure of human type III 3alpha-hydroxysteroid dehydrogenase (HSD)/bile acid binding protein (AKR1C2) complexed with NADP(+) and 3alpha,7beta-dihydroxy-5beta-cholanic acid (ursodeoxycholate) at 3.0 A resolution is presented. Thus, the three-dimensional structure has now been solved for a human HSD member of the aldo-keto reductase superfamily. AKR1C2 is implicated in the prostatic production of the potent androgen 5alpha-dihydrotestosterone and the hepatic transport of bile acids. It also catalyzes the formation of the neurosteroid 3alpha-hydroxy-5alpha-pregnan-20-one in the central nervous system, and its allosteric modulation by fluoxetine has been linked to the use of this drug for premenstrual dsyphoria. Like other members of the superfamily, AKR1C2 folds into an alpha/beta-barrel and binds NADP(+) in an extended conformation. The carboxylate of ursodeoxycholate binds to AKR1C2 in the oxyanion hole at the active site. More interestingly, the orientation of ursodeoxycholate is essentially "backwards" and "upside-down" from that observed for testosterone in the related rat 3alpha-HSD.NADP(+).testosterone ternary complex, where testosterone assumes the position of a 3-ketosteroid substrate. The orientation of ursodeoxycholate is thus similar to that expected of a 17beta-HSD substrate. The ternary structure explains the ability of AKR1C2 to catalyze 3alpha-, 17beta-, and 20alpha-HSD reactions. Comparison of the steroid binding pocket of AKR1C2 with that of rat 3alpha-HSD reveals significant differences in the positions of conserved and nonconserved loop residues, providing insights into the structural basis for the functional flexibility that is observed in all the human 3alpha-HSD isoforms but not in the rat isoform.
- Department of Pharmacology, Department of Biochemistry and Biophysics, and The Johnson Research Foundation, University of Pennsylvania School of Medicine, Philadelphia, Pennsylvania 19104, USA.
Organizational Affiliation: