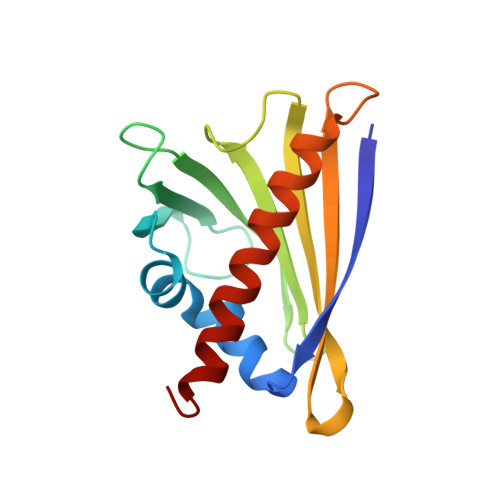
Crystal structures of two homologous pathogenesis-related proteins from yellow lupine.
Biesiadka, J., Bujacz, G., Sikorski, M.M., Jaskolski, M.(2002) J Mol Biology 319: 1223-1234
- PubMed: 12079359 Search on PubMed
- DOI: https://doi.org/10.1016/S0022-2836(02)00385-6
- Primary Citation Related Structures:
1ICX, 1IFV - PubMed Abstract:
Pathogenesis-related class 10 (PR10) proteins are restricted to the plant kingdom where they are coded by multigene families and occur at high levels. In spite of their abundance, their physiological role is obscure although members of a distantly related subclass (cytokinin-specific binding proteins) are known to bind plant hormones. PR10 proteins are of special significance in legume plants where their expression patterns are related to infection by the symbiotic, nitrogen-fixing bacteria. Here we present the first crystal structures of classic PR10 proteins representing two homologues from one subclass in yellow lupine. The general fold is similar and, as in a birch pollen allergen, consists of a seven-stranded beta-sheet wrapped around a long C-terminal helix. The mouth of a large pocket formed between the beta-sheet and the helix seems a likely site for ligand binding. The shape of the pocket varies because, in variance with the rigid beta-sheet, the helix shows unusual conformational variability consisting in bending, disorder, and axial shifting. A surface loop, proximal to the entrance to the internal cavity, shows an unusual structural conservation and rigidity in contrast to the high glycine content in its sequence. The loop is different from the so-called glycine-rich P-loops that bind phosphate groups of nucleotides, but it is very likely that it does play a role in ligand binding in PR10 proteins.
- Institute of Bioorganic Chemistry, Polish Academy of Sciences, 61-704 Poznan, Poland.
Organizational Affiliation: